
Coral Reefs Optimization (CRO)
Contents
Introduction
Bioinspired algorithms based on natural processes and systems have recently attracted considerable interest from developers. Coral Reef Optimization (CRO) algorithm, originally proposed by S. Salcedo-Sanz et al. in 2014, is probably the most notable among them.
The CRO algorithm is based on modeling the processes of formation and development of coral reefs in nature. These processes include various mechanisms of coral reproduction (broadcast spawning, brooding, as well as asexual reproduction), competition for limited space in the reef, and the death of weak individuals. Just as evolution creates resilient and adaptable coral reefs in nature, the CRO algorithm allows one to explore the search space and find optimal or near-optimal solutions to various problems.
In this paper, we present an improved version of the CROm algorithm with a modified destruction mechanism based on the use of the inverse power law distribution to generate new solutions in the neighborhood of the best ones. The proposed approach not only preserves the traditional advantages of CRO, such as exploratory capacity and a natural balance between global exploration and local exploitation of the search space, but also complements them with a more efficient mechanism that allows for more accurate localization of promising search areas and faster convergence to optimal solutions.
We are going to extensively test the proposed algorithm on a set of classical optimization benchmark functions demonstrating its improved performance compared to the original CRO algorithm and other modern metaheuristics. The experimental results show that the proposed approach is particularly effective for problems with multimodal objective functions and complex search landscape structure.
The article is structured as follows: first, we review the basic concepts and operating principles of the standard CRO algorithm, then we describe in detail the proposed modifications and their theoretical justification. This is followed by an experimental evaluation of the algorithm and analysis of the results. In conclusion, we discuss the obtained results, limitations of the proposed method, and possible directions for future research.
Implementation of the algorithm
The basic idea of CRO is to simulate coral colonies that develop and compete for space on a reef, ultimately forming an optimal structure. Each coral in the reef represents a potential solution to the optimization problem under consideration.
The reef is modeled as a two-dimensional N×M grid. Each grid cell can either be occupied by a coral or left empty. A coral is a coded solution to an optimization problem. For each coral, a fitness (health) function is determined that corresponds to the objective function of the optimization problem.
The ρ₀ ∈ (0,1) parameter determines the initial proportion of the reef occupied by corals, that is, the ratio of occupied cells to the total number of cells at the beginning of the algorithm. Initialization of the reef is performed as follows:
- The reef size of N×M and the initial filling fraction ρ₀ are specified.
- ⌊ρ₀ × N × M⌋ reef cells are randomly selected to house the starting corals.
- Initial corals are generated randomly within the search area and placed in the selected cells.
After the reef is initialized, an iterative process of reef formation and development begins, consisting of several stages:
Broadcast Spawning. For this type of reproduction, a certain proportion of Fₑ existing corals is selected. The selected corals form pairs and create offspring using crossover operators. Each pair produces a larva using the crossover operator (ordinary averaging).
Brooding. The remaining fraction of corals (1-Fₑ) engage in brooding where each coral produces offspring through mutation. For each coral selected for brooding, a larva is created using a mutation operator. The larva typically represents a small random variation of the encoded solution.
Larval settlement. After the larvae are formed, each one tries to take its place in the reef during the reproductive stages. The settlement is carried out according to the following rules:
- The larva randomly chooses a cell (i, j) in the reef.
- If the cell is free, the larva occupies it.
- If the cell is occupied, the larva can displace the existing coral only if its fitness is higher: f(larva) > f(Ξᵢⱼ).
- If displacement does not occur, the larva may try to settle in another place (up to a maximum k number of attempts).
- If after k attempts the larva fails to find a place, it dies.
Asexual reproduction (budding). The best corals in a reef (Fₐ fraction) can reproduce asexually, creating exact copies of themselves (clones). Formally:
- Corals are sorted by the fitness function.
- The best Fₐ × 100% corals are selected for asexual reproduction.
- Each selected coral creates a clone that attempts to settle in the reef according to the same rules as during the larval settlement.
Depredation. At the end of each iteration, the worst corals in the reef may die with Pd probability, making room for new corals in the next iterations.
The reef formation is repeated until the specified stopping criterion is met, such as reaching the maximum number of iterations. After stopping, the best coral in the reef represents the found solution to the optimization problem.
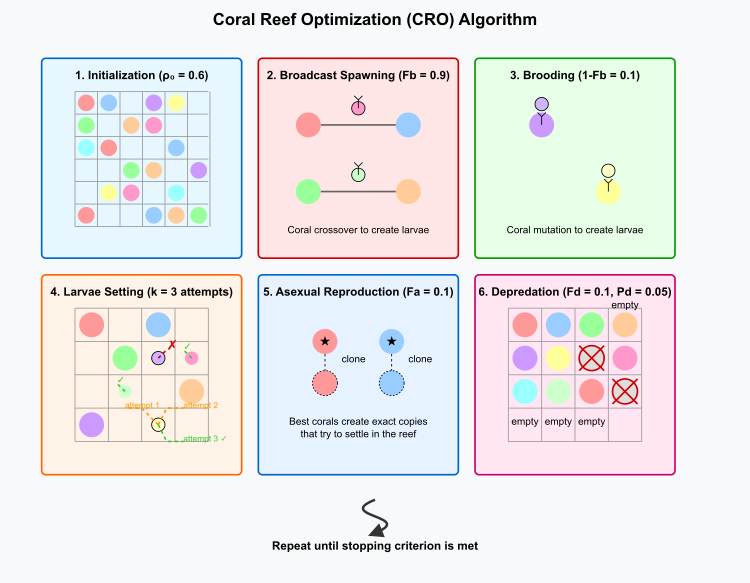
Figure 1. CRO algorithm works as follows:
The image above shows all six main stages of the algorithm:
- Initialization (ρ₀ = 0.6) displays a two-dimensional grid (reef) partially filled with multi-colored corals representing different solutions.
- Broadcast spawning (Fb = 0.9) shows corals forming pairs and creating larvae through crossover.
- Brooding (1-Fb = 0.1) demonstrates individual corals producing larvae through mutation.
- Larval settlement (k = 3 attempts) illustrates the larval search for a place in the reef, including successful and unsuccessful attempts.
- Asexual reproduction (Fa = 0.1) shows the best corals (marked with stars) that create exact copies of themselves.
- Depredation (Fd = 0.1, Pd = 0.05) demonstrates the removal of the worst corals from the reef.
Input: Reef parameters (N, M, ρ₀, Fₑ, Fₐ, Fd, Pd, k)
Output: Best solution found
1. Initialization:
- Create a N×M reef
- Fill ⌊ρ₀ × N × M⌋ cells with random corals
- Calculate the fitness of each coral
2. Until the stopping criterion is met:
3. Broadcast spawning:
- Select a share of Fₑ corals to participate
- Form pairs and create larvae through crossover
4. Brooding:
- For the remaining corals (1-Fₑ), create larvae through mutation
5. Larval settlement:
- For each larva:
- Try to occupy a random reef cell
- If occupied, displace existing coral with higher fitness
- If unsuccessful, try again (up to k attempts)
6. Asexual reproduction:
- Sort corals by fitness
- The best Fₐ corals create clones
- Clones attempt to settle in the reef
7. Depredation:
- With Pd probability to perform destruction
- Remove the Fd share of the worst corals
8. Return the best coral to the reef
Now we can code the CRO algorithm. Let's implement the class called C_AO_CRO, which implements the CRO algorithm and inherits from the C_AO base class.
CRO parameters:
The class declares parameters that control the algorithm behavior:
- popSize — coral population size.
- reefRows — reef height (number of rows in the grid).
- reefCols — reef height (number of columns in the grid).
- rho0 — initial reef occupancy. This is the percentage of reef cells occupied by corals at the start of the algorithm.
- Fb — proportion of corals participating in broadcast spawning.
- Fa — proportion of corals participating in asexual reproduction (fragmentation).
- Fd — proportion of corals subject to removal due to low fitness.
- Pd — probability of coral destruction.
- attemptsNum — number of attempts the larva makes to settle on the reef.
Class methods:
- SetParams () — method sets the values of the algorithm parameters based on the values stored in the "params" array. This is an array that allows us to dynamically change the algorithm parameters during execution.
- Init () — initialization method that accepts search ranges for optimization variables (rangeMinP, rangeMaxP, rangeStepP) and the number of epochs (epochsP). The Init method performs the necessary preparations to start the algorithm, such as initializing the population and reef.
- Moving () — basic logic of corals moving along the reef.
- Revision () — revise and adjust the positions of corals on the reef.
- InitReef () — initialize the reef structure preparing it for colonization by corals.
- LarvaSettling () — determine where the coral larva will settle on the reef, simulating reef colonization by new corals. The "larva" parameter represents the coral larva.
- BroadcastSpawning () — corals release larvae into the water.
- Brooding () — larvae develop inside the coral.
- AsexualReproduction () — corals fragment and form new colonies.
- Depredation () — the corals are destroyed.
- GetReefCoordIndex () — convert reef coordinates (row and column) to an index in a one-dimensional array.
- SortAgentsByFitness () — sort corals according to their fitness.
Private variables:
- totalReefSize — total size of the reef (product of reefRows and reefCols).
- occupied [] — array of boolean values indicating whether each reef cell is occupied by a coral.
- reefIndices [] — array of indices containing the indices of corals (from the a [] array of the C_AO base class) in occupied reef cells.
//—————————————————————————————————————————————————————————————————————————————— class C_AO_CRO : public C_AO { public: //-------------------------------------------------------------------- ~C_AO_CRO () { } C_AO_CRO () { ao_name = "CRO"; ao_desc = "Coral Reef Optimization"; ao_link = "https://www.mql5.com/en/articles/17760"; popSize = 50; // population size reefRows = 20; // reef height reefCols = 20; // reef width rho0 = 0.2; // initial reef occupancy Fb = 0.99; // fraction of corals for broadcast spawning Fa = 0.01; // fraction of corals for asexual reproduction Fd = 0.8; // fraction of corals to remove Pd = 0.9; // probability of destruction attemptsNum = 20; // number of attempts for larvae to settle ArrayResize (params, 9); params [0].name = "popSize"; params [0].val = popSize; params [1].name = "reefRows"; params [1].val = reefRows; params [2].name = "reefCols"; params [2].val = reefCols; params [3].name = "rho0"; params [3].val = rho0; params [4].name = "Fb"; params [4].val = Fb; params [5].name = "Fa"; params [5].val = Fa; params [6].name = "Fd"; params [6].val = Fd; params [7].name = "Pd"; params [7].val = Pd; params [8].name = "attemptsNum"; params [8].val = attemptsNum; } void SetParams () { popSize = (int)params [0].val; reefRows = (int)params [1].val; reefCols = (int)params [2].val; rho0 = params [3].val; Fb = params [4].val; Fa = params [5].val; Fd = params [6].val; Pd = params [7].val; attemptsNum = (int)params [8].val; } bool Init (const double &rangeMinP [], // minimum search range const double &rangeMaxP [], // maximum search range const double &rangeStepP [], // search step const int epochsP = 0); // number of epochs void Moving (); void Revision (); //---------------------------------------------------------------------------- int reefRows; // reef height int reefCols; // reef width double rho0; // initial reef occupancy double Fb; // fraction of corals for broadcast spawning double Fa; // fraction of corals for asexual reproduction double Fd; // fraction of corals to remove double Pd; // probability of destruction int attemptsNum; // number of attempts for larvae to settle private: //------------------------------------------------------------------- int totalReefSize; // total reef size bool occupied []; // flags of reef cell occupancy int reefIndices []; // agent indices in a[] corresponding to occupied cells // Auxiliary methods void InitReef (); int LarvaSettling (S_AO_Agent &larva); void BroadcastSpawning (S_AO_Agent &larvae [], int &larvaCount); void Brooding (S_AO_Agent &larvae [], int &larvaCount); void AsexualReproduction (); void Depredation (); int GetReefCoordIndex (int row, int col); void SortAgentsByFitness (int &indices [], int &count); }; //——————————————————————————————————————————————————————————————————————————————
The Init method initializes the CRO algorithm: it calls the StandardInit of the base class, then calculates the totalReefSize, determines the initial number of corals (initialPopSize), initializes the "occupied" and the "reefIndices" array, calls InitReef() to place corals on the reef, and finally returns 'true' on success.
//—————————————————————————————————————————————————————————————————————————————— bool C_AO_CRO::Init (const double &rangeMinP [], // minimum search range const double &rangeMaxP [], // maximum search range const double &rangeStepP [], // search step const int epochsP = 0) // number of epochs { // Standard initialization of the parent class if (!StandardInit (rangeMinP, rangeMaxP, rangeStepP)) return false; //---------------------------------------------------------------------------- // Calculate the reef total size totalReefSize = reefRows * reefCols; // The number of starting corals should not exceed popSize int initialPopSize = (int)MathRound (rho0 * totalReefSize); if (initialPopSize > popSize) initialPopSize = popSize; // Initialize the occupancy array and indices ArrayResize (occupied, totalReefSize); ArrayResize (reefIndices, totalReefSize); // Fill the arrays with initial values for (int i = 0; i < totalReefSize; i++) { occupied [i] = false; reefIndices [i] = -1; } // Reef initialization InitReef (); return true; } //——————————————————————————————————————————————————————————————————————————————The InitReef() method of the C_AO_CRO class initializes the initial population of corals on the reef. First, we calculate the number of corals to initialize based on the given density (rho0) of the reef. This number is limited by the popSize population size. Then, for each coral, a random but free position on the reef is allocated, once it is found, it is marked as occupied, and the coral is associated with this position in the reefIndices array. Next, coordinates are generated for each coral. Each coordinate is selected randomly within the given boundaries (rangeMin, rangeMax) and discretized using the u.SeInDiSp function, which takes into account the discretization step (rangeStep). This ensures a uniform and controlled distribution of initial solutions (corals) in the search space.
//—————————————————————————————————————————————————————————————————————————————— void C_AO_CRO::InitReef () { // Number of starting corals in the reef (based on rho0) int initialCorals = (int)MathRound (rho0 * totalReefSize); // The number of starting corals should not exceed the population size if (initialCorals > popSize) initialCorals = popSize; // Initialize initialCorals random positions in the reef for (int i = 0; i < initialCorals; i++) { int pos; // Look for a free position do { pos = (int)MathFloor (u.RNDfromCI (0, totalReefSize)); // Protection against exceeding the array size if (pos < 0) pos = 0; if (pos >= totalReefSize) pos = totalReefSize - 1; } while (occupied [pos]); // Create a new coral at the found position occupied [pos] = true; reefIndices [pos] = i; // Generate random coordinates for a new coral for (int c = 0; c < coords; c++) { double coordinate = u.RNDfromCI (rangeMin [c], rangeMax [c]); a [i].c [c] = u.SeInDiSp (coordinate, rangeMin [c], rangeMax [c], rangeStep [c]); } } } //——————————————————————————————————————————————————————————————————————————————
The Moving() method performs an initial estimate of the coral population. If "revision" is false, the method iterates through all reef positions. For each occupied position (determined by the value of the "occupied" array element), the method gets the index of the coral located at that position from the reefIndices array. If the resulting idx index is valid, the fitness of this coral will be calculated.
It is important to note that the Moving() method does not directly calculate the fitness itself, but only provides the necessary information to calculate it. After iterating over all reef positions, the method sets the "revision" value to 'true' to prevent the coral evaluation loop from running again in the same optimization iteration. Essentially, the Moving() method acts as a trigger for calculating the coral fitness, ensuring that this process is performed once per optimization iteration.
//—————————————————————————————————————————————————————————————————————————————— void C_AO_CRO::Moving () { if (!revision) { // Initial assessment of all corals in the reef for (int i = 0; i < totalReefSize; i++) { if (occupied [i]) { int idx = reefIndices [i]; if (idx >= 0 && idx < popSize) { // Calculating fitness does not require using GetFitness() // since it will be evaluated in external code (in FuncTests) } } } revision = true; } } //——————————————————————————————————————————————————————————————————————————————
The Revision() method updates the global best solution. It searches for corals on the reef with fitness better than the current global best solution and replaces it. It creates an array, preparing a place for the larvae produced through reproduction. The method then initiates sexual reproduction - it calls the BroadcastSpawning and Brooding methods to create larvae. The next step of the method involves the larval settlement involving the LarvaSettling method to determine locations for new larvae on the reef. AsexualReproduction is initiated, followed by destruction. In short, it updates the solution, spawns corals, deploys larvae, and simulates the impact of depredation.
//—————————————————————————————————————————————————————————————————————————————— void C_AO_CRO::Revision () { // Update the global best solution for (int i = 0; i < totalReefSize; i++) { if (occupied [i] && a [reefIndices [i]].f > fB) { fB = a [reefIndices [i]].f; ArrayCopy (cB, a [reefIndices [i]].c, 0, 0, WHOLE_ARRAY); } } // Form an array to store larvae S_AO_Agent larvae []; ArrayResize (larvae, totalReefSize * 2); // Allocate with reserve int larvaCount = 0; // Stage 1: Broadcast Spawning BroadcastSpawning (larvae, larvaCount); // Stage 2: Brooding Brooding (larvae, larvaCount); // Calculate the fitness function for each larva // (will be executed in external code in FuncTests) // Stage 3: Larval settlement for (int i = 0; i < larvaCount; i++) { LarvaSettling (larvae [i]); } // Stage 4: Asexual reproduction AsexualReproduction (); // Stage 5: Depredation Depredation (); } //——————————————————————————————————————————————————————————————————————————————
The LarvaSettling method simulates the process of larval settlement on a reef. It tries to find a suitable place for the larva and either settles it in a vacant place or replaces an existing, but less fit coral. When attempting to settle, the larva attempts to settle attemptsNum times. On each attempt, a random position on the reef is selected. If the chosen position is free (not occupied by a coral), the method searches for a free index in the a[] agents' array, if a free index is found, the larva copies its solution (c[] coordinates and f fitness) into the corresponding cell of the agents' array.
The information that the reef position is now occupied is updated, and the method returns the index of the successfully occupied 'pos' position. If the chosen position is occupied, the method compares the fitness of the larva with the fitness of the current coral in that position. If the larva is better, it replaces the existing coral by copying its parameters to its cell in the agents array, and the method returns the "pos" index of the position the replacement occurred at. If after all attempts the larva fails to settle (either in an unoccupied space or by replacing an existing coral), the method returns -1.
//—————————————————————————————————————————————————————————————————————————————— int C_AO_CRO::LarvaSettling (S_AO_Agent &larva) { // Try to settle the larva attemptsNum times for (int attempt = 0; attempt < attemptsNum; attempt++) { // Select a random position in the reef int pos = (int)MathFloor (u.RNDfromCI (0, totalReefSize)); // Check that the position is within the array if (pos < 0 || pos >= totalReefSize) continue; // If the position is free, populate the larva if (!occupied [pos]) { // Search for a free index in the agents array int newIndex = -1; for (int i = 0; i < popSize; i++) { bool used = false; for (int j = 0; j < totalReefSize; j++) { if (reefIndices [j] == i) { used = true; break; } } if (!used) { newIndex = i; break; } } if (newIndex != -1) { // Copy the larva's solution ArrayCopy (a [newIndex].c, larva.c, 0, 0, WHOLE_ARRAY); a [newIndex].f = larva.f; // Update information about the reef occupied [pos] = true; reefIndices [pos] = newIndex; return pos; } } // If the position is occupied, check if the larva is better than the current coral else if (occupied [pos] && reefIndices [pos] >= 0 && reefIndices [pos] < popSize && larva.f > a [reefIndices [pos]].f) { // The larva displaces the existing coral ArrayCopy (a [reefIndices [pos]].c, larva.c, 0, 0, WHOLE_ARRAY); a [reefIndices [pos]].f = larva.f; return pos; } } // If the larva failed to settle, return -1 return -1; } //——————————————————————————————————————————————————————————————————————————————
The BroadcastSpawning method simulates the broadcast spawning. It performs the following basic steps: it finds the indices of all coral-occupied sites on the reef, determines the number of corals participating in spawning based on the Fb (Brooding Fraction) parameter - the proportion of corals participating in broadcast spawning, mixes the indices of occupied corals and then selects pairs for spawning. Creation of larvae by crossover: : for each pair of corals, a new larva is created, the coordinates of the larva are calculated as the average value of the coordinates of the parents with the addition of a small random mutation. The crossover aims to combine the best traits of the parents, and the resulting larvae are added to the "larvae" array.
Key points:
- Fb here is responsible for the fraction of corals in the broadcast spawning, while in Brooding it is responsible for the fraction of corals engaged in brooding gestation.
- Crossover and mutation increase the genetic diversity of a population.
- The aim of the method is to create new larvae obtained from a combination of genes of parent corals for further colonization of the reef. The method involves paired crossover.
//—————————————————————————————————————————————————————————————————————————————— void C_AO_CRO::BroadcastSpawning (S_AO_Agent &larvae [], int &larvaCount) { // Find all occupied positions int occupiedIndices []; int occupiedCount = 0; for (int i = 0; i < totalReefSize; i++) { if (occupied [i]) { ArrayResize (occupiedIndices, occupiedCount + 1); occupiedIndices [occupiedCount] = i; occupiedCount++; } } // Check if there are no occupied positions if (occupiedCount == 0) return; // Select the Fb share for broadcast spawning int broadcastCount = (int)MathRound (Fb * occupiedCount); if (broadcastCount <= 0) broadcastCount = 1; // At least one coral if (broadcastCount > occupiedCount) broadcastCount = occupiedCount; // Shuffle the indices for (int i = 0; i < occupiedCount; i++) { // Register the array out-of-bounds problem int j = (int)MathFloor (u.RNDfromCI (0, occupiedCount)); // Ensure that j is within the array bounds if (j >= 0 && j < occupiedCount && i < occupiedCount) { int temp = occupiedIndices [i]; occupiedIndices [i] = occupiedIndices [j]; occupiedIndices [j] = temp; } } // Form pairs and create offspring for (int i = 0; i < broadcastCount - 1; i += 2) { if (i + 1 < broadcastCount) // Make sure there is a second parent { int idx1 = reefIndices [occupiedIndices [i]]; int idx2 = reefIndices [occupiedIndices [i + 1]]; if (idx1 >= 0 && idx1 < popSize && idx2 >= 0 && idx2 < popSize) { // Initialize the larva larvae [larvaCount].Init (coords); // Create a new larva as a result of crossover for (int c = 0; c < coords; c++) { // Simple crossover method: average of parents' coordinates with a small mutation double value = (a [idx1].c [c] + a [idx2].c [c]) / 2.0 + u.RNDfromCI (-0.1, 0.1) * (rangeMax [c] - rangeMin [c]); larvae [larvaCount].c [c] = u.SeInDiSp (value, rangeMin [c], rangeMax [c], rangeStep [c]); } // Increase the larvae counter larvaCount++; } } } } //——————————————————————————————————————————————————————————————————————————————
The Brooding method simulates the internal larval gestation of corals in CRO and follows these steps:
- determines, which locations on the reef are occupied by corals, and stores indices of these positions.
- calculates how many corals will participate in brooding using the Fb (Brooding Fraction) parameter. The indices of the occupied positions are shuffled for a randomly selected corals participating in reproduction.
- creates and mutate larvae: for each selected coral, a larva is created. The larva initializes and mutates: its coordinates deviate slightly from those of the parent coral at random. Mutated larvae are added to the "larvae" array.
Key points:
- Fb defines the proportion of corals that do NOT engage in asexual reproduction (i.e., engage in internal gestation).
- The mutation provides gene diversity in the larval population.
- The goal is to create new, potentially superior individuals that will compete for space on the reef.
//—————————————————————————————————————————————————————————————————————————————— void C_AO_CRO::Brooding (S_AO_Agent &larvae [], int &larvaCount) { // Find all occupied positions int occupiedIndices []; int occupiedCount = 0; for (int i = 0; i < totalReefSize; i++) { if (occupied [i]) { ArrayResize (occupiedIndices, occupiedCount + 1); occupiedIndices [occupiedCount] = i; occupiedCount++; } } // Check if there are no occupied positions if (occupiedCount == 0) return; // Number of corals for internal reproduction int broodingCount = (int)MathRound ((1.0 - Fb) * occupiedCount); if (broodingCount <= 0) broodingCount = 1; // At least one coral if (broodingCount > occupiedCount) broodingCount = occupiedCount; // Shuffle the indices for (int i = 0; i < occupiedCount; i++) { // Register the array out-of-bounds problem int j = (int)MathFloor (u.RNDfromCI (0, occupiedCount)); // Ensure that j is within the array bounds if (j >= 0 && j < occupiedCount && i < occupiedCount) { int temp = occupiedIndices [i]; occupiedIndices [i] = occupiedIndices [j]; occupiedIndices [j] = temp; } } // For each selected coral, create a mutated copy for (int i = 0; i < broodingCount; i++) { if (i < occupiedCount) // Check for out-of-bounds { int idx = reefIndices [occupiedIndices [i]]; if (idx >= 0 && idx < popSize) { // Initialize the larva larvae [larvaCount].Init (coords); // Create a new larva as a mutation of the original coral for (int c = 0; c < coords; c++) { // Mutation: add a small random deviation double value = a [idx].c [c] + u.RNDfromCI (-0.2, 0.2) * (rangeMax [c] - rangeMin [c]); larvae [larvaCount].c [c] = u.SeInDiSp (value, rangeMin [c], rangeMax [c], rangeStep [c]); } // Increase the larvae counter larvaCount++; } } } } //——————————————————————————————————————————————————————————————————————————————
The AsexualReproduction method simulates asexual reproduction of corals by budding. The method performs the following steps: it finds the indices of all the positions occupied by corals on the reef, sorts the indices of the occupied positions in order of decreasing fitness, that is, the positions with the fittest corals come first. Based on the "Fa" (Fraction for Asexual reproduction) parameter, which specifies the proportion of corals participating in asexual reproduction, it defines the number of the best corals that will produce clones. Creation and colonization of clones: for each selected coral, an exact copy (clone) is created — a larva with the same coordinates and adaptations. The LarvaSettling method is called to attempt to settle the clone on the reef.
Key points:
- Asexual reproduction allows the most adaptable corals to quickly spread their genes.
- The Fa parameter controls the intensity of asexual reproduction.
- Using LarvaSettling to populate clones means that the clones should compete for space on the reef in the same way as sexually produced larvae.
- The cloning method, in this case, is an exact copy of the parent WITHOUT mutations. Mutations occur only during sexual reproduction.
//—————————————————————————————————————————————————————————————————————————————— void C_AO_CRO::AsexualReproduction () { // Find all occupied positions and their indices int occupiedIndices []; int occupiedCount = 0; for (int i = 0; i < totalReefSize; i++) { if (occupied [i]) { ArrayResize (occupiedIndices, occupiedCount + 1); occupiedIndices [occupiedCount] = i; occupiedCount++; } } // If there are no occupied positions, exit if (occupiedCount == 0) return; // Sort indices by fitness SortAgentsByFitness (occupiedIndices, occupiedCount); // Select the best Fa% of corals for asexual reproduction int budCount = (int)MathRound (Fa * occupiedCount); if (budCount <= 0) budCount = 1; // At least one coral if (budCount > occupiedCount) budCount = occupiedCount; // For each selected coral, create a clone and try to populate it for (int i = 0; i < budCount; i++) { if (i < occupiedCount) // Check for out-of-bounds { int idx = reefIndices [occupiedIndices [i]]; if (idx >= 0 && idx < popSize) { // Create a new larva as an exact copy of the original coral S_AO_Agent clone; clone.Init (coords); ArrayCopy (clone.c, a [idx].c, 0, 0, WHOLE_ARRAY); clone.f = a [idx].f; // Try to populate the clone LarvaSettling (clone); } } } } //——————————————————————————————————————————————————————————————————————————————
The Depredation method simulates the coral extinction due to predation or other negative factors (e.g. pollution). In this case, destruction is applied with Pd probability. If the random number is less than Pd, the destruction is performed. The search for occupied positions is determined by the indices of all reef positions occupied by corals. The method sorts the indices of occupied positions by coral fitness.
The array of sorted indices is reversed so that the indices of the least fit corals are at the beginning of the array. The amount of corals to be removed is determined based on the Fd (Fraction for Depredation) parameter. This value is rounded to the nearest integer. The specified number of least adapted corals are removed. Next comes the reef cleaning: for each position scheduled for deletion: the corresponding cell in the "occupied" array is set to 'false' indicating that the position is now free. The corresponding cell in the reefIndices array is set to -1, indicating that there is no coral at that position.
//—————————————————————————————————————————————————————————————————————————————— oid C_AO_CRO::Depredation() { // Apply destruction with Pd probability if (u.RNDfromCI(0, 1) < Pd) { // Find all occupied positions and their indices int occupiedIndices[]; int occupiedCount = 0; for (int i = 0; i < totalReefSize; i++) { if (occupied[i]) { ArrayResize(occupiedIndices, occupiedCount + 1); occupiedIndices[occupiedCount] = i; occupiedCount++; } } // If there are no occupied positions, exit if (occupiedCount == 0) return; // Sort indices by fitness SortAgentsByFitness(occupiedIndices, occupiedCount); // Reverse the array so the worst ones are first for (int i = 0; i < occupiedCount / 2; i++) { if(i < occupiedCount && (occupiedCount - 1 - i) < occupiedCount) // Check for out-of-bounds { int temp = occupiedIndices[i]; occupiedIndices[i] = occupiedIndices[occupiedCount - 1 - i]; occupiedIndices[occupiedCount - 1 - i] = temp; } } // Remove the worst Fd% corals int removeCount = (int)MathRound(Fd * occupiedCount); if (removeCount <= 0) removeCount = 1; // At least one coral if (removeCount > occupiedCount) removeCount = occupiedCount; // Overflow protection for (int i = 0; i < removeCount; i++) { if(i < occupiedCount) // Check for out-of-bounds { int pos = occupiedIndices[i]; if(pos >= 0 && pos < totalReefSize) // Additional check { occupied[pos] = false; reefIndices[pos] = -1; } } } } } //——————————————————————————————————————————————————————————————————————————————
The GetReefCoordIndex method converts reef coordinates (row and column) into a one-dimensional index and takes "row" and "col" as inputs, returning the corresponding index in the array representing the reef. First, the method checks whether the passed coordinates are outside the boundaries of the reef. Otherwise, it calculates the index using the equation (row * reefCols + col, where "reefCols" is the number of columns in the reef). The method is used to access elements in arrays representing the reef.
//—————————————————————————————————————————————————————————————————————————————— int C_AO_CRO::GetReefCoordIndex (int row, int col) { // Check for out-of-bounds if (row < 0 || row >= reefRows || col < 0 || col >= reefCols) return -1; // Convert a two-dimensional position to a one-dimensional index return row * reefCols + col; } //——————————————————————————————————————————————————————————————————————————————
The SortAgentsByFitness method sorts the "indices" array of "count" elements in descending order of their fitness using a bubble sort algorithm based on the fitness values stored in the "a" array accessed through the reefIndices array. Thus, after running the "indices" method, it contains coral indices sorted from the fittest to the least fit. Additional checks are included to prevent array bounds being exceeded.
//—————————————————————————————————————————————————————————————————————————————— void C_AO_CRO::SortAgentsByFitness (int &indices [], int &count) { // Check for an empty array if (count <= 0) return; // Bubble sort by decreasing fitness for (int i = 0; i < count - 1; i++) { for (int j = 0; j < count - i - 1; j++) { if (j + 1 < count) // Check that j+1 is not out of bounds { int idx1 = reefIndices [indices [j]]; int idx2 = reefIndices [indices [j + 1]]; if (idx1 >= 0 && idx1 < popSize && idx2 >= 0 && idx2 < popSize) // Check indices { if (a [idx1].f < a [idx2].f) { int temp = indices [j]; indices [j] = indices [j + 1]; indices [j + 1] = temp; } } } } } } //——————————————————————————————————————————————————————————————————————————————
We have finished preparing the algorithm and described all the numerous methods. Now we can proceed directly to testing.
Test results
After conducting tests, it is clear that the CRO algorithm is working rather poorly and it is necessary to reconsider some of the methods of the original approach.CRO|Coral Reef Optimization|50.0|5.0|5.0|0.4|0.9|0.1|0.1|0.01|3.0|
=============================
5 Hilly's; Func runs: 10000; result: 0.365266682511984
25 Hilly's; Func runs: 10000; result: 0.270828009448956
500 Hilly's; Func runs: 10000; result: 0.2504192846772352
=============================
5 Forest's; Func runs: 10000; result: 0.23618879234608753
25 Forest's; Func runs: 10000; result: 0.19453106526100442
500 Forest's; Func runs: 10000; result: 0.1679109693993047
=============================
5 Megacity's; Func runs: 10000; result: 0.13076923076923075
25 Megacity's; Func runs: 10000; result: 0.11138461538461542
500 Megacity's; Func runs: 10000; result: 0.09366153846153921
=============================
All score: 1.82096 (20.23%)
The first thing I noticed about the CRO algorithm was the destruction method. It needs to be modified to improve the search. We will replace the worst corals on the reef with new ones generated near the best corals (solutions), thereby pushing the search toward more promising areas. We will determine the number of the best (eliteCount) and the worst (removeCount) corals.
Replacing the worst solutions: For each "prey," an "elite" coral is selected, and a new coral is generated in the vicinity of this "elite" coral. In the improved destruction mechanism, an inverse power-law distribution with exponent 10 (where 'power' = 0.1) is applied to generate deviations from the best solution. This distribution is characterized by the fact that most new solutions are generated in the immediate vicinity of the best one, but with a small probability, significant deviations may occur, which ensures a balance between local search (exploitation) and global exploration of the solution space. The new coordinates are limited by the allowed range. The vacated position on the reef is then populated by a new coral.
//—————————————————————————————————————————————————————————————————————————————— void C_AO_CRO::Depredation () { // Apply destruction with Pd probability if (u.RNDfromCI (0, 1) < Pd) { // Find all occupied positions and their indices int occupiedIndices []; int occupiedCount = 0; for (int i = 0; i < totalReefSize; i++) { if (occupied [i]) { ArrayResize (occupiedIndices, occupiedCount + 1); occupiedIndices [occupiedCount] = i; occupiedCount++; } } // If there are no occupied positions, exit if (occupiedCount == 0) return; // Sort the indices by fitness (from best to worst) SortAgentsByFitness (occupiedIndices, occupiedCount); // Determine the number of best solutions used to generate new ones int eliteCount = (int)MathRound (0.1 * occupiedCount); // Use top 10% if (eliteCount <= 0) eliteCount = 1; // At least one elite solution if (eliteCount > occupiedCount) eliteCount = occupiedCount; // Determine the number of worst solutions to be replaced int removeCount = (int)MathRound (Fd * occupiedCount); if (removeCount <= 0) removeCount = 1; // At least one solution is replaced if (removeCount > occupiedCount - eliteCount) removeCount = occupiedCount - eliteCount; // Do not remove elite solutions // Remove the worst solutions and replace them with new ones in the vicinity of the best ones for (int i = 0; i < removeCount; i++) { // Index of the solution to be removed (from the end of the sorted array) int removeIndex = occupiedCount - 1 - i; if (removeIndex < 0 || removeIndex >= occupiedCount) continue; int posToRemove = occupiedIndices [removeIndex]; if (posToRemove < 0 || posToRemove >= totalReefSize) continue; // Choose one of the elite solutions double power = 0.1; // Power distribution parameter double r = u.RNDfromCI (0, 1); int eliteIdx = (int)(eliteCount); if (eliteIdx >= eliteCount) eliteIdx = eliteCount - 1; int posBest = occupiedIndices [eliteIdx]; if (posBest < 0 || posBest >= totalReefSize) continue; int bestAgentIdx = reefIndices [posBest]; if (bestAgentIdx < 0 || bestAgentIdx >= popSize) continue; // Free up space for a new solution occupied [posToRemove] = false; // Generate a new solution in the neighborhood of the selected elite solution int newAgentIdx = reefIndices [posToRemove]; // Use the same agent index if (newAgentIdx >= 0 && newAgentIdx < popSize) { // Generation via power-law distribution around the best solution for (int c = 0; c < coords; c++) { // Determine the search radius (can be adapted depending on progress) double radius = 0.7 * (rangeMax [c] - rangeMin [c]); // 10% of the range // Generate a value using a power law double sign = u.RNDfromCI (0, 1) < 0.5 ? -1.0 : 1.0; // Random sign double deviation = sign * radius * MathPow (u.RNDfromCI (0, 1), 1.0 / power); // New value in the neighborhood of the best one double newValue = a [bestAgentIdx].c [c] + deviation; // Limit the value to an acceptable range a [newAgentIdx].c [c] = u.SeInDiSp (newValue, rangeMin [c], rangeMax [c], rangeStep [c]); } // Populate the new solution into the reef occupied [posToRemove] = true; reefIndices [posToRemove] = newAgentIdx; } } } } //——————————————————————————————————————————————————————————————————————————————
Now, after the changes have been made, let's test the CROm algorithm. As you can see from the results below, the algorithm has improved its performance significantly.
CROm|Coral Reef Optimization M|50.0|20.0|20.0|0.2|0.99|0.01|0.8|0.9|20.0|
=============================
5 Hilly's; Func runs: 10000; result: 0.7851210159578113
25 Hilly's; Func runs: 10000; result: 0.4603296933002806
500 Hilly's; Func runs: 10000; result: 0.25958379129490083
=============================
5 Forest's; Func runs: 10000; result: 0.8668751980437325
25 Forest's; Func runs: 10000; result: 0.3529695710837671
500 Forest's; Func runs: 10000; result: 0.16267582083006701
=============================
5 Megacity's; Func runs: 10000; result: 0.6323076923076923
25 Megacity's; Func runs: 10000; result: 0.2673846153846154
500 Megacity's; Func runs: 10000; result: 0.10733846153846247
=============================
All score: 3.89459 (43.27%)
The algorithm's visualization is not particularly noteworthy. The algorithm has a slightly more difficult time coping with discrete tasks on the Megacity test function.
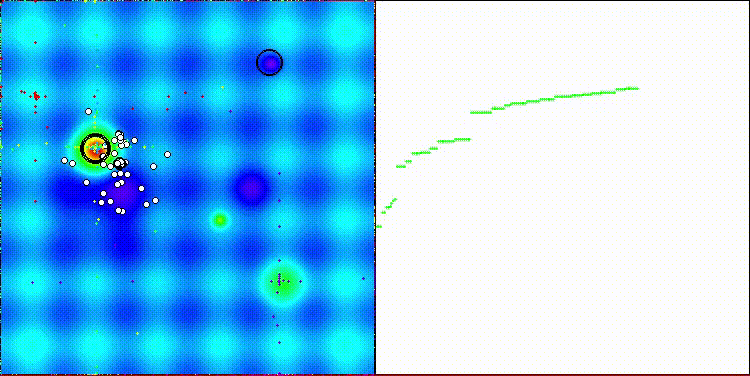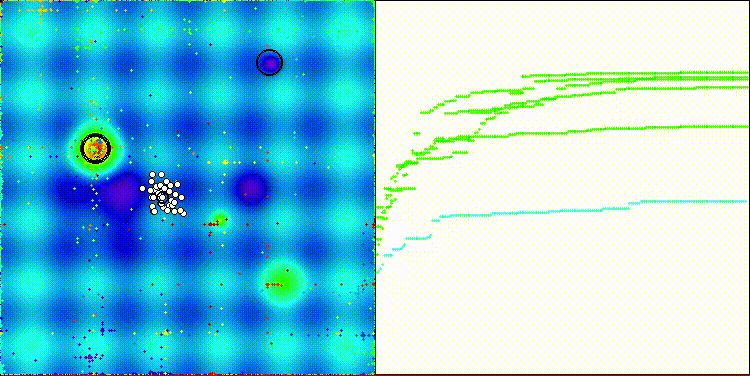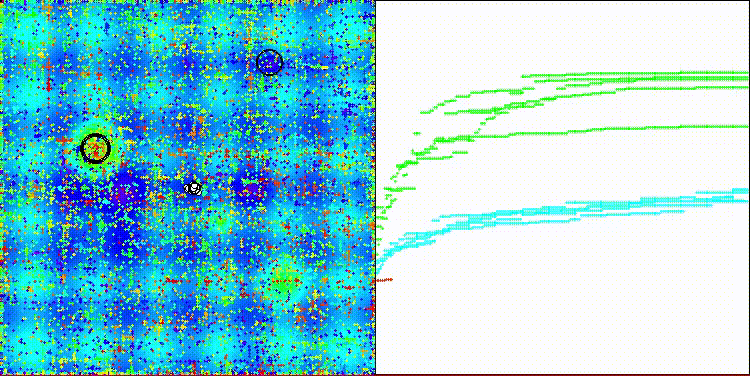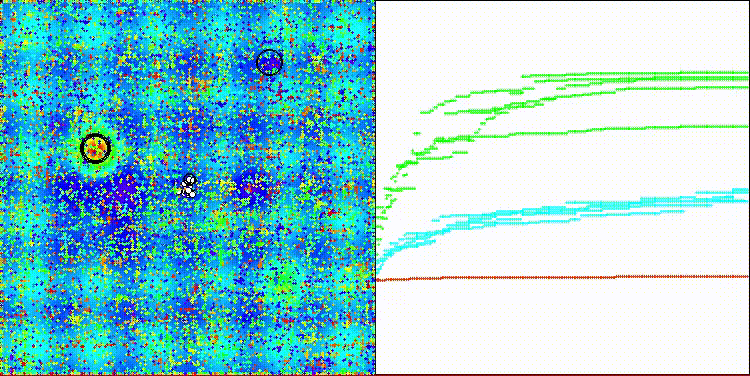
CROm on the Hilly test function
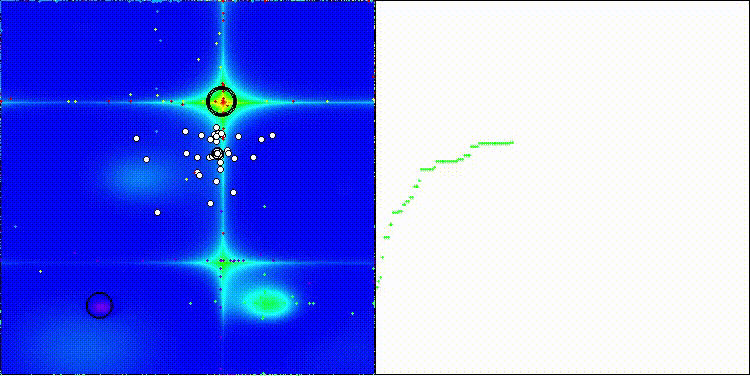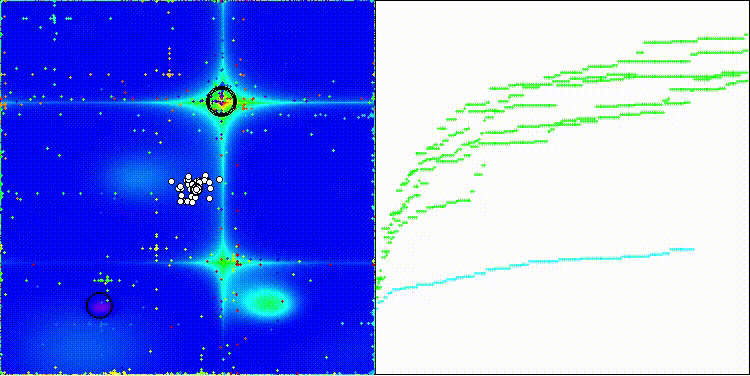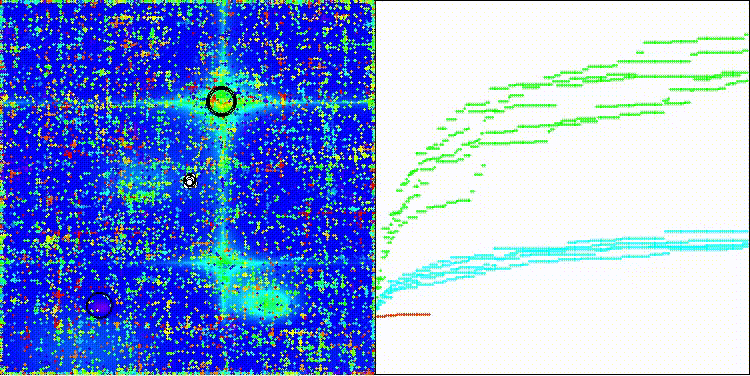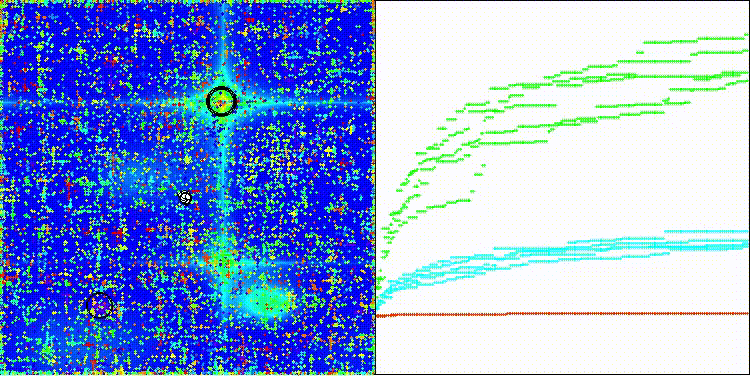
CROm on the Forest test function
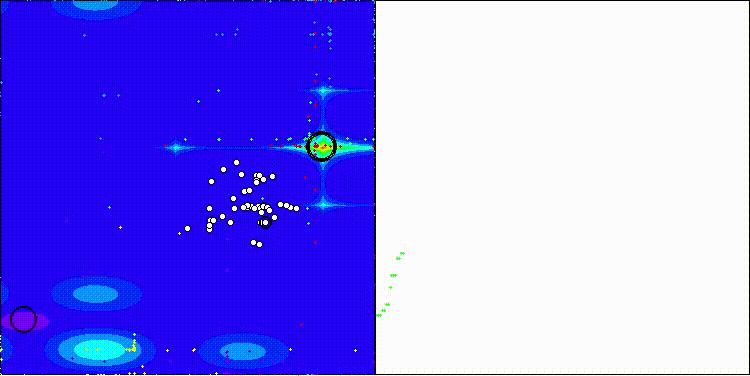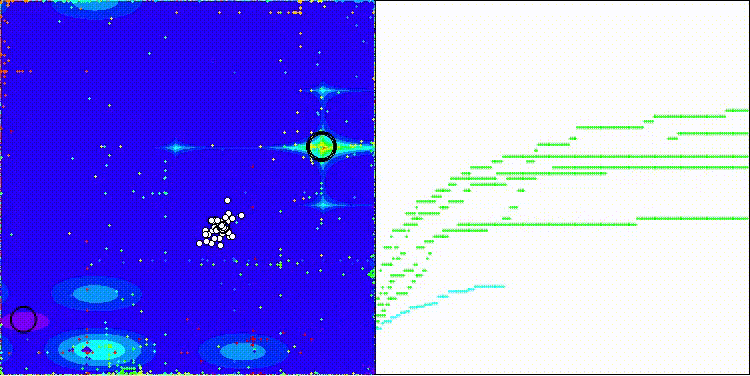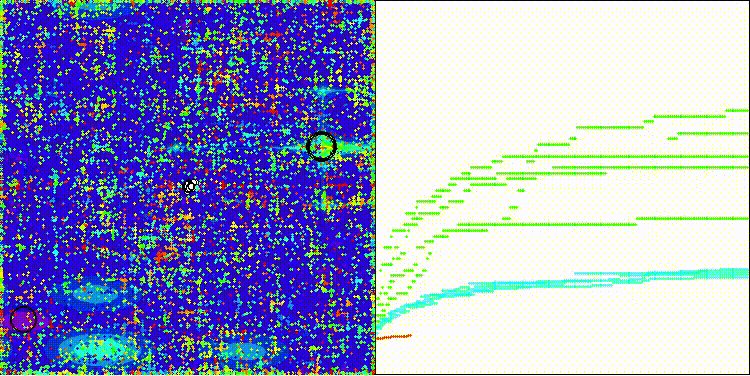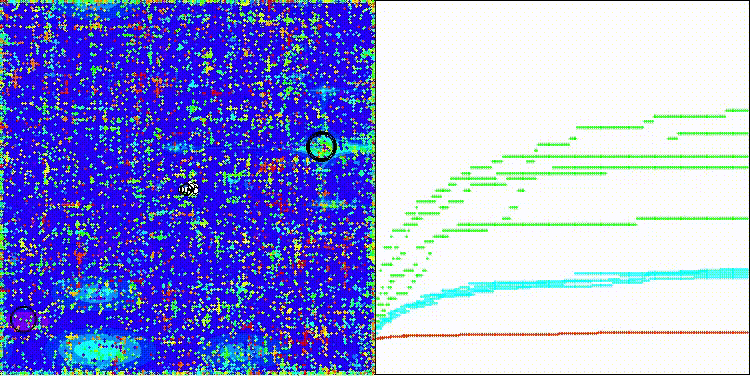
CROm on the Megacity test function
Based on the test results, the CROm algorithm is ranked 42nd in the ranking table of population-based optimization algorithms.
| # | AO | Description | Hilly | Hilly Final | Forest | Forest Final | Megacity (discrete) | Megacity Final | Final Result | % of MAX | ||||||
| 10 p (5 F) | 50 p (25 F) | 1000 p (500 F) | 10 p (5 F) | 50 p (25 F) | 1000 p (500 F) | 10 p (5 F) | 50 p (25 F) | 1000 p (500 F) | ||||||||
| 1 | ANS | across neighbourhood search | 0.94948 | 0.84776 | 0.43857 | 2.23581 | 1.00000 | 0.92334 | 0.39988 | 2.32323 | 0.70923 | 0.63477 | 0.23091 | 1.57491 | 6.134 | 68.15 |
| 2 | CLA | code lock algorithm (joo) | 0.95345 | 0.87107 | 0.37590 | 2.20042 | 0.98942 | 0.91709 | 0.31642 | 2.22294 | 0.79692 | 0.69385 | 0.19303 | 1.68380 | 6.107 | 67.86 |
| 3 | AMOm | animal migration ptimization M | 0.90358 | 0.84317 | 0.46284 | 2.20959 | 0.99001 | 0.92436 | 0.46598 | 2.38034 | 0.56769 | 0.59132 | 0.23773 | 1.39675 | 5.987 | 66.52 |
| 4 | (P+O)ES | (P+O) evolution strategies | 0.92256 | 0.88101 | 0.40021 | 2.20379 | 0.97750 | 0.87490 | 0.31945 | 2.17185 | 0.67385 | 0.62985 | 0.18634 | 1.49003 | 5.866 | 65.17 |
| 5 | CTA | comet tail algorithm (joo) | 0.95346 | 0.86319 | 0.27770 | 2.09435 | 0.99794 | 0.85740 | 0.33949 | 2.19484 | 0.88769 | 0.56431 | 0.10512 | 1.55712 | 5.846 | 64.96 |
| 6 | TETA | time evolution travel algorithm (joo) | 0.91362 | 0.82349 | 0.31990 | 2.05701 | 0.97096 | 0.89532 | 0.29324 | 2.15952 | 0.73462 | 0.68569 | 0.16021 | 1.58052 | 5.797 | 64.41 |
| 7 | SDSm | stochastic diffusion search M | 0.93066 | 0.85445 | 0.39476 | 2.17988 | 0.99983 | 0.89244 | 0.19619 | 2.08846 | 0.72333 | 0.61100 | 0.10670 | 1.44103 | 5.709 | 63.44 |
| 8 | BOAm | billiards optimization algorithm M | 0.95757 | 0.82599 | 0.25235 | 2.03590 | 1.00000 | 0.90036 | 0.30502 | 2.20538 | 0.73538 | 0.52523 | 0.09563 | 1.35625 | 5.598 | 62.19 |
| 9 | AAm | archery algorithm M | 0.91744 | 0.70876 | 0.42160 | 2.04780 | 0.92527 | 0.75802 | 0.35328 | 2.03657 | 0.67385 | 0.55200 | 0.23738 | 1.46323 | 5.548 | 61.64 |
| 10 | ESG | evolution of social groups (joo) | 0.99906 | 0.79654 | 0.35056 | 2.14616 | 1.00000 | 0.82863 | 0.13102 | 1.95965 | 0.82333 | 0.55300 | 0.04725 | 1.42358 | 5.529 | 61.44 |
| 11 | SIA | simulated isotropic annealing (joo) | 0.95784 | 0.84264 | 0.41465 | 2.21513 | 0.98239 | 0.79586 | 0.20507 | 1.98332 | 0.68667 | 0.49300 | 0.09053 | 1.27020 | 5.469 | 60.76 |
| 12 | ACS | artificial cooperative search | 0.75547 | 0.74744 | 0.30407 | 1.80698 | 1.00000 | 0.88861 | 0.22413 | 2.11274 | 0.69077 | 0.48185 | 0.13322 | 1.30583 | 5.226 | 58.06 |
| 13 | DA | dialectical algorithm | 0.86183 | 0.70033 | 0.33724 | 1.89940 | 0.98163 | 0.72772 | 0.28718 | 1.99653 | 0.70308 | 0.45292 | 0.16367 | 1.31967 | 5.216 | 57.95 |
| 14 | BHAm | black hole algorithm M | 0.75236 | 0.76675 | 0.34583 | 1.86493 | 0.93593 | 0.80152 | 0.27177 | 2.00923 | 0.65077 | 0.51646 | 0.15472 | 1.32195 | 5.196 | 57.73 |
| 15 | ASO | anarchy society optimization | 0.84872 | 0.74646 | 0.31465 | 1.90983 | 0.96148 | 0.79150 | 0.23803 | 1.99101 | 0.57077 | 0.54062 | 0.16614 | 1.27752 | 5.178 | 57.54 |
| 16 | RFO | royal flush optimization (joo) | 0.83361 | 0.73742 | 0.34629 | 1.91733 | 0.89424 | 0.73824 | 0.24098 | 1.87346 | 0.63154 | 0.50292 | 0.16421 | 1.29867 | 5.089 | 56.55 |
| 17 | AOSm | atomic orbital search M | 0.80232 | 0.70449 | 0.31021 | 1.81702 | 0.85660 | 0.69451 | 0.21996 | 1.77107 | 0.74615 | 0.52862 | 0.14358 | 1.41835 | 5.006 | 55.63 |
| 18 | TSEA | turtle shell evolution algorithm (joo) | 0.96798 | 0.64480 | 0.29672 | 1.90949 | 0.99449 | 0.61981 | 0.22708 | 1.84139 | 0.69077 | 0.42646 | 0.13598 | 1.25322 | 5.004 | 55.60 |
| 19 | DE | differential evolution | 0.95044 | 0.61674 | 0.30308 | 1.87026 | 0.95317 | 0.78896 | 0.16652 | 1.90865 | 0.78667 | 0.36033 | 0.02953 | 1.17653 | 4.955 | 55.06 |
| 20 | SRA | successful restaurateur algorithm (joo) | 0.96883 | 0.63455 | 0.29217 | 1.89555 | 0.94637 | 0.55506 | 0.19124 | 1.69267 | 0.74923 | 0.44031 | 0.12526 | 1.31480 | 4.903 | 54.48 |
| 21 | CRO | chemical reaction optimization | 0.94629 | 0.66112 | 0.29853 | 1.90593 | 0.87906 | 0.58422 | 0.21146 | 1.67473 | 0.75846 | 0.42646 | 0.12686 | 1.31178 | 4.892 | 54.36 |
| 22 | BIO | blood inheritance optimization (joo) | 0.81568 | 0.65336 | 0.30877 | 1.77781 | 0.89937 | 0.65319 | 0.21760 | 1.77016 | 0.67846 | 0.47631 | 0.13902 | 1.29378 | 4.842 | 53.80 |
| 23 | BSA | bird swarm algorithm | 0.89306 | 0.64900 | 0.26250 | 1.80455 | 0.92420 | 0.71121 | 0.24939 | 1.88479 | 0.69385 | 0.32615 | 0.10012 | 1.12012 | 4.809 | 53.44 |
| 24 | HS | harmony search | 0.86509 | 0.68782 | 0.32527 | 1.87818 | 0.99999 | 0.68002 | 0.09590 | 1.77592 | 0.62000 | 0.42267 | 0.05458 | 1.09725 | 4.751 | 52.79 |
| 25 | SSG | saplings sowing and growing | 0.77839 | 0.64925 | 0.39543 | 1.82308 | 0.85973 | 0.62467 | 0.17429 | 1.65869 | 0.64667 | 0.44133 | 0.10598 | 1.19398 | 4.676 | 51.95 |
| 26 | BCOm | bacterial chemotaxis optimization M | 0.75953 | 0.62268 | 0.31483 | 1.69704 | 0.89378 | 0.61339 | 0.22542 | 1.73259 | 0.65385 | 0.42092 | 0.14435 | 1.21912 | 4.649 | 51.65 |
| 27 | ABO | african buffalo optimization | 0.83337 | 0.62247 | 0.29964 | 1.75548 | 0.92170 | 0.58618 | 0.19723 | 1.70511 | 0.61000 | 0.43154 | 0.13225 | 1.17378 | 4.634 | 51.49 |
| 28 | (PO)ES | (PO) evolution strategies | 0.79025 | 0.62647 | 0.42935 | 1.84606 | 0.87616 | 0.60943 | 0.19591 | 1.68151 | 0.59000 | 0.37933 | 0.11322 | 1.08255 | 4.610 | 51.22 |
| 29 | TSm | tabu search M | 0.87795 | 0.61431 | 0.29104 | 1.78330 | 0.92885 | 0.51844 | 0.19054 | 1.63783 | 0.61077 | 0.38215 | 0.12157 | 1.11449 | 4.536 | 50.40 |
| 30 | BSO | brain storm optimization | 0.93736 | 0.57616 | 0.29688 | 1.81041 | 0.93131 | 0.55866 | 0.23537 | 1.72534 | 0.55231 | 0.29077 | 0.11914 | 0.96222 | 4.498 | 49.98 |
| 31 | WOAm | wale optimization algorithm M | 0.84521 | 0.56298 | 0.26263 | 1.67081 | 0.93100 | 0.52278 | 0.16365 | 1.61743 | 0.66308 | 0.41138 | 0.11357 | 1.18803 | 4.476 | 49.74 |
| 32 | AEFA | artificial electric field algorithm | 0.87700 | 0.61753 | 0.25235 | 1.74688 | 0.92729 | 0.72698 | 0.18064 | 1.83490 | 0.66615 | 0.11631 | 0.09508 | 0.87754 | 4.459 | 49.55 |
| 33 | AEO | artificial ecosystem-based optimization algorithm | 0.91380 | 0.46713 | 0.26470 | 1.64563 | 0.90223 | 0.43705 | 0.21400 | 1.55327 | 0.66154 | 0.30800 | 0.28563 | 1.25517 | 4.454 | 49.49 |
| 34 | ACOm | ant colony optimization M | 0.88190 | 0.66127 | 0.30377 | 1.84693 | 0.85873 | 0.58680 | 0.15051 | 1.59604 | 0.59667 | 0.37333 | 0.02472 | 0.99472 | 4.438 | 49.31 |
| 35 | BFO-GA | bacterial foraging optimization - ga | 0.89150 | 0.55111 | 0.31529 | 1.75790 | 0.96982 | 0.39612 | 0.06305 | 1.42899 | 0.72667 | 0.27500 | 0.03525 | 1.03692 | 4.224 | 46.93 |
| 36 | SOA | simple optimization algorithm | 0.91520 | 0.46976 | 0.27089 | 1.65585 | 0.89675 | 0.37401 | 0.16984 | 1.44060 | 0.69538 | 0.28031 | 0.10852 | 1.08422 | 4.181 | 46.45 |
| 37 | ABHA | artificial bee hive algorithm | 0.84131 | 0.54227 | 0.26304 | 1.64663 | 0.87858 | 0.47779 | 0.17181 | 1.52818 | 0.50923 | 0.33877 | 0.10397 | 0.95197 | 4.127 | 45.85 |
| 38 | ACMO | atmospheric cloud model optimization | 0.90321 | 0.48546 | 0.30403 | 1.69270 | 0.80268 | 0.37857 | 0.19178 | 1.37303 | 0.62308 | 0.24400 | 0.10795 | 0.97503 | 4.041 | 44.90 |
| 39 | ADAMm | adaptive moment estimation M | 0.88635 | 0.44766 | 0.26613 | 1.60014 | 0.84497 | 0.38493 | 0.16889 | 1.39880 | 0.66154 | 0.27046 | 0.10594 | 1.03794 | 4.037 | 44.85 |
| 40 | CGO | chaos game optimization | 0.57256 | 0.37158 | 0.32018 | 1.26432 | 0.61176 | 0.61931 | 0.62161 | 1.85267 | 0.37538 | 0.21923 | 0.19028 | 0.78490 | 3.902 | 43.35 |
| 41 | ATAm | artificial tribe algorithm M | 0.71771 | 0.55304 | 0.25235 | 1.52310 | 0.82491 | 0.55904 | 0.20473 | 1.58867 | 0.44000 | 0.18615 | 0.09411 | 0.72026 | 3.832 | 42.58 |
| 42 | CROm | coral reefs optimization M | 0.78512 | 0.46032 | 0.25958 | 1.50502 | 0.86688 | 0.35297 | 0.16267 | 1.38252 | 0.63231 | 0.26738 | 0.10734 | 1.00703 | 3.895 | 43.27 |
| 43 | CFO | central force optimization | 0.60961 | 0.54958 | 0.27831 | 1.43750 | 0.63418 | 0.46833 | 0.22541 | 1.32792 | 0.57231 | 0.23477 | 0.09586 | 0.90294 | 3.668 | 40.76 |
| 44 | ASHA | artificial showering algorithm | 0.89686 | 0.40433 | 0.25617 | 1.55737 | 0.80360 | 0.35526 | 0.19160 | 1.35046 | 0.47692 | 0.18123 | 0.09774 | 0.75589 | 3.664 | 40.71 |
| 45 | ASBO | adaptive social behavior optimization | 0.76331 | 0.49253 | 0.32619 | 1.58202 | 0.79546 | 0.40035 | 0.26097 | 1.45677 | 0.26462 | 0.17169 | 0.18200 | 0.61831 | 3.657 | 40.63 |
| RW | neuroboids optimization algorithm 2(joo) | 0.48754 | 0.32159 | 0.25781 | 1.06694 | 0.37554 | 0.21944 | 0.15877 | 0.75375 | 0.27969 | 0.14917 | 0.09847 | 0.52734 | 2.348 | 26.09 | |
Summary
Living coral reefs are finely tuned systems that continually maintain a balance between stability and variability, between the preservation of proven survival strategies and the experimentation with new ones. A modified depredation mechanism, with the generation of new solutions in the vicinity of the best ones, mimics the natural process of ecological success in coral reefs.
In nature, after the death of some colonies, it is not random species that appear in the vacated areas of the reef, but those that have most successfully adapted to local environmental conditions. Moreover, the survival rate of new colonies is higher in zones surrounding successful existing colonies, which is directly reflected in the algorithm by generating new solutions around elite ones. This ensures the constant exploration of promising areas of the search space and maintains a constant population size.
Implementing an inverse power law distribution (with exponent 10, where power = 0.1) allows generating deviations from the best solution. This distribution concentrates most new solutions close to the best with rare significant deviations, providing an optimal balance between local exploitation and global exploration.
Increasing the search radius to 70% of the feasible range allows the algorithm to explore wider areas of the solution space.
Using only the best solutions (elite) as a basis for generating new solutions accelerates convergence to optimal regions and improves the quality of the resulting solutions, while a variety of operators simulating key aspects of coral evolution: broadcast spawning and brooding, larval settlement, and asexual reproduction can be used in the development of other population-based optimization methods.
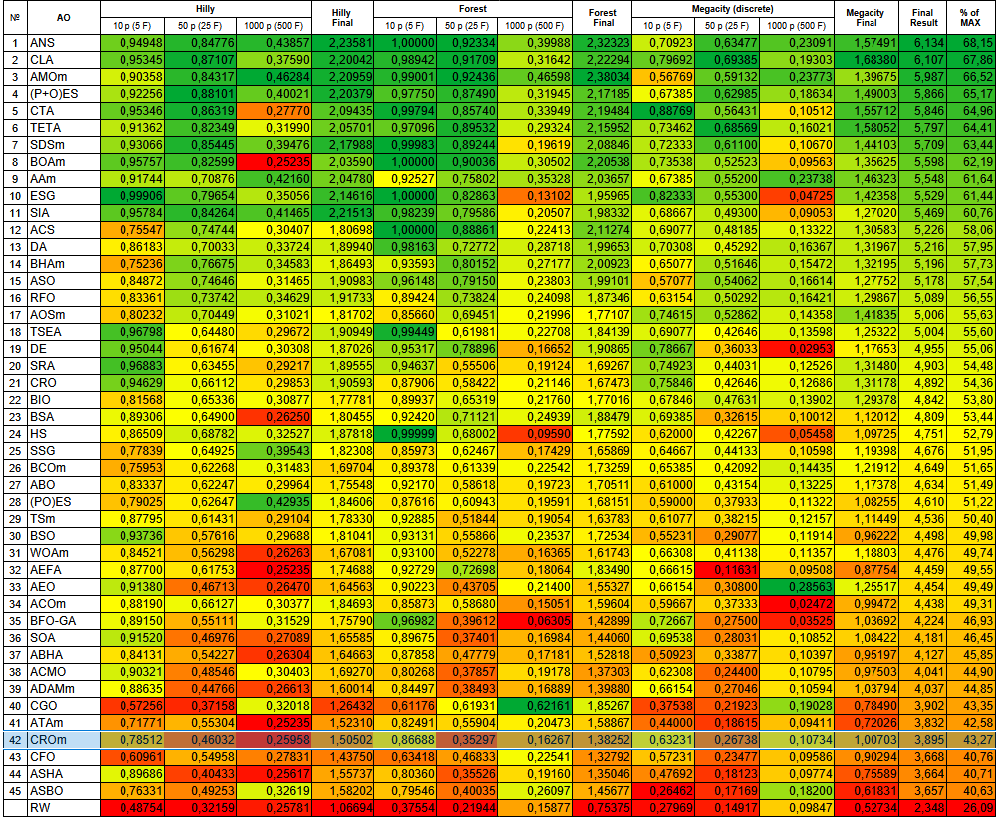
Figure 2. Color gradation of algorithms according to the corresponding tests
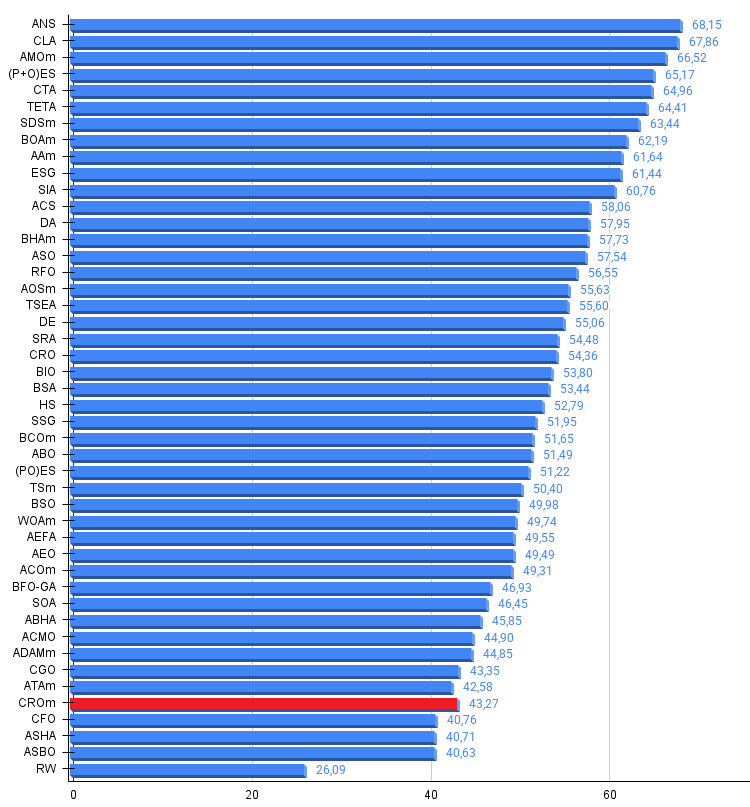
Figure 3. Histogram of algorithm testing results (scale from 0 to 100, the higher the better, where 100 is the maximum possible theoretical result, in the archive there is a script for calculating the rating table)
CROm pros and cons:
Pros:
- Interesting idea.
- Promising development.
- Stable results.
Cons:
- Weaker results on discrete functions.
- A large number of external parameters.
The article is accompanied by an archive with the current versions of the algorithm codes. The author of the article is not responsible for the absolute accuracy in the description of canonical algorithms. Changes have been made to many of them to improve search capabilities. The conclusions and judgments presented in the articles are based on the results of the experiments.
Programs used in the article
| # | Name | Type | Description |
|---|---|---|---|
| 1 | #C_AO.mqh | Include | Parent class of population optimization algorithms |
| 2 | #C_AO_enum.mqh | Include | Enumeration of population optimization algorithms |
| 3 | TestFunctions.mqh | Include | Library of test functions |
| 4 | TestStandFunctions.mqh | Include | Test stand function library |
| 5 | Utilities.mqh | Include | Library of auxiliary functions |
| 6 | CalculationTestResults.mqh | Include | Script for calculating results in the comparison table |
| 7 | Testing AOs.mq5 | Script | The unified test stand for all population optimization algorithms |
| 8 | Simple use of population optimization algorithms.mq5 | Script | A simple example of using population optimization algorithms without visualization |
| 9 | Test_AO_CROm.mq5 | Script | CROm test stand |
Translated from Russian by MetaQuotes Ltd.
Original article: https://www.mql5.com/ru/articles/17760
Warning: All rights to these materials are reserved by MetaQuotes Ltd. Copying or reprinting of these materials in whole or in part is prohibited.
This article was written by a user of the site and reflects their personal views. MetaQuotes Ltd is not responsible for the accuracy of the information presented, nor for any consequences resulting from the use of the solutions, strategies or recommendations described.
 Price Action Analysis Toolkit Development (Part 65): Building an MQL5 System to Monitor and Analyze Manually Drawn Fibonacci Levels
Price Action Analysis Toolkit Development (Part 65): Building an MQL5 System to Monitor and Analyze Manually Drawn Fibonacci Levels
 Creating Custom Indicators in MQL5 (Part 10): Enhancing the Footprint Chart with Per-Bar Volume Sentiment Information Box
Creating Custom Indicators in MQL5 (Part 10): Enhancing the Footprint Chart with Per-Bar Volume Sentiment Information Box
 From Novice to Expert: Adaptive Risk Management for Liquidity Strategies
From Novice to Expert: Adaptive Risk Management for Liquidity Strategies
 Developing Market Entropy Indicator: Trading System Based on Information Theory
Developing Market Entropy Indicator: Trading System Based on Information Theory
- Free trading apps
- Over 8,000 signals for copying
- Economic news for exploring financial markets
You agree to website policy and terms of use