
Biogeography-Based Optimization (BBO)
Contents
Introduction
While reviewing some optimization algorithms, I became interested in the Biogeography-Based Optimization (BBO) algorithm, which was developed by Professor Dan Simon in 2008. BBO draws inspiration from biogeography, the study of the geographical distribution of biological organisms. Mathematical models describing species distribution patterns were first developed in the 1960s. Just as genetic algorithms were inspired by biological genetics and neural networks by biological neurons, BBO uses the mathematical principles of biogeography to solve optimization problems.
In nature, islands of an archipelago with favorable conditions (high Habitat Suitability Index - HSI) have a large number of species and high emigration, while islands with poor conditions have few species and high immigration. This natural dynamic of species migration between islands formed the basis of the BBO optimization mechanism. The algorithm uses the concept of species migration to exchange characteristics between solutions, the mutation probability is based on a theoretically sound species distribution model, and good solutions actively share their characteristics but remain robust to change. This feature is one of the algorithm’s defining characteristics.
In this article, we examine this elegant algorithmic concept, implement it in code, and evaluate the performance of the BBO algorithm.
Implementation of the algorithm
Imagine archipelagos of islands, where each island is home to different species of animals.
1. Habitat = Island = Problem solution. Each island in our algorithm represents one possible solution. If you have 50 islands, then you have 50 different solutions.
2. HSI (Habitat Suitability Index) = Island suitability for living = Solution quality. Rich island with fresh water, fruits and good climate = Good solution (high HSI). Desert island with no water = Bad solution (low HSI)
3. Species = Solution characteristics. A resource-rich island hosts many species, whereas a poor island hosts few because its habitat is less diverse.
How does migration work? Real-life example: Hawaii (rich island), many species → animals often swim/fly away to other islands (high emigration), but few swim to the island (low immigration - the island is already overcrowded). Uninhabited island: few species → animals rarely leave (low emigration), but new ones often arrive (high immigration - lots of free space).
In the algorithm: Bad solution (few species) → High immigration → takes on characteristics of good solutions. Good solution (many types) → High emigration → shares characteristics with others.
Another example from life: let's say we are looking for the best location for a store in the city. Each "island" is a location option. Create 50 random locations (islands), of which:
Island 1: Bad place - outskirts
Island 2: Great location - center
Island 50: Average good place
Rate each location (HSI): Island 2 (center): HSI = 95 (many shoppers, good accessibility), Island 1 (outskirts): HSI = 20 (few shoppers), then Island 1 (bad) has high immigration and it "accepts" some characteristics from Island 2 (good). For example, if Island 2 is located "near a subway station", then Island 1 will also try to find a place near a subway station. Sometimes "disasters" (earthquakes, tsunamis) can occur, when the solution changes by accident — the store "jumps" to a completely new location, and this movement helps find unexpectedly good solutions.
Why do average solutions mutate less often? In nature, very rich islands (like the Galapagos) are rare and unstable, while very poor islands are also rare and unstable, while average islands are the most common and stable. In the algorithm this means:
Very good solutions (HSI = 95): high probability of mutation
Very bad solutions (HSI = 5): high probability of mutation
Average solutions (HSI = 50): low probability of mutation
The first 2 best islands (solutions) are protected from changes - these are our "reserves". We do not want to lose the best solutions we have found! Then the final optimization process looks as follows: we create 50 random solutions (islands), sort them by quality (from best to worst), then bad solutions learn from good ones, and some solutions are randomly changed. In this way, BBO simulates the natural process of species distribution between islands to find an optimal solution. Below is an illustration of how the algorithm works.
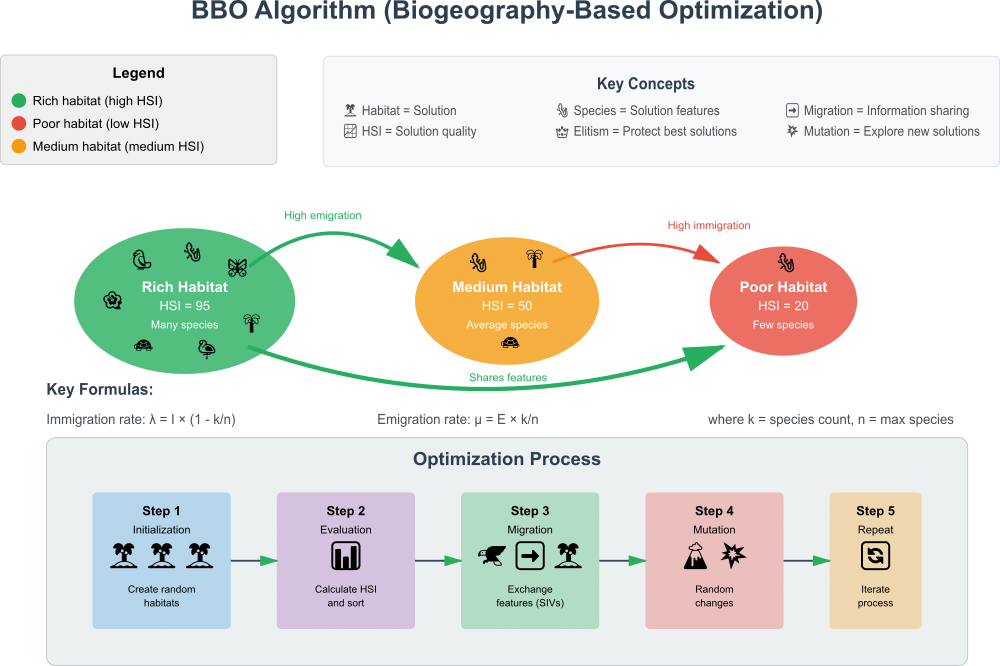
Figure 1. BBO algorithm in action
The diagram shows:
- Three types of islands — (rich, medium, poor) with different number of species
- Migration — arrows show how species move between islands
- Step-by-step optimization — from initialization to repetition
- Key concepts — legend and basic algorithm principles
The illustration clearly demonstrates how rich islands (good solutions) have high emigration, poor islands (bad solutions) have high immigration, there is an exchange of characteristics between solutions, and the entire optimization cycle works. Let's prepare a pseudo code.
1. INITIALIZATION:
- Set the parameters:
* population size (number of habitats) = 50
* maximum immigration rate I = 1.0
* maximum emigration rate E = 1.0
* mutation probability = 0.01
* number of elite solutions = 2
* maximum number of species = 50
* number of iterations
- Create a population of N random habitats (solutions)
- Calculate the HSI (suitability) for each habitat
- Calculate the probabilities of existence for different numbers of species
2. MAIN LOOP (repeat a specified number of iterations):
2.1. EVALUATION AND SORTING:
- Calculate the HSI for each habitat
- Sort habitats by descending HSI
- Save the best solution
2.2. CALCULATION OF MIGRATION PARAMETERS:
For each i habitat:
- Determine the number of species: Si = Smax × (N - rank_i) / N
- Calculate the immigration rate: λi = I × (1 - Si/Smax)
- Calculate the emigration rate: μi = E × (Si/Smax)
- Determine the probability of existence based on Si
2.3. MIGRATION (exchange of characteristics):
For each Hi habitat (except elite):
IF random_number < λi (immigration rate), THEN:
For each decision variable (SIV) j:
IF random_number < λi, THEN:
- Select a donor habitat:
* calculate the sum of all emigration rates (except Hi)
* use roulette selection based on μ
* select habitat Hk with the probability of μk/Σμ
- Copy the j-th variable from Hk to Hi
END IF
END of the loop on variables
END IF
END of the cycle on habitats
2.4. MUTATION (exploration of new solutions):
For each Hi habitat (except elite):
- Calculate the mutation rate: m_rate = m × (1 - probability_of_existence_i)
IF random_number < m_rate, THEN:
- Select a random variable j
- Replace it with a random value from the acceptable range
END IF
END of the cycle on habitats
2.5. REPLACEMENT AND UPDATE:
- Calculate new HSI values
- Update the best solution found
- Save current fitness values for the next iteration
3. RETURN the best solution found
Now we only need to implement the C_AO_BBO class, which will be derived from the C_AO class and will be designed to implement the BBO algorithm. Inheritance implies that C_AO_BBO uses the base functionality for optimization provided by the parent class.
The class contains a set of parameters, population size and BBO-specific ones, such as: maximum immigration/emigration rate (immigrationMax, emigrationMax), mutation probability (mutationProb), number of elite solutions kept unchanged (elitismCount), and maximum number of species (speciesMax). The class constructor initializes the BBO parameters with default values, assigns a name, description, and a link to an article about the algorithm. The SetParams() method allows changing the values of parameters using data from the "params" array.
Main methods:- Init () — initializes the algorithm, including creating and initializing the population, specifying the ranges of values, the step and number of epochs, and initializing the arrays to store the habitat data.
- Moving () — implements the basic logic of moving (migrating) solutions between habitats in accordance with BBO principles.
- Revision () — includes logic for revising and adjusting solutions (habitats).
- S_HabitatData — internal structure containing information about each habitat (solution), including the number of species (speciesCount), immigration/emigration rate (immigration, emigration), and probability of existence (probability).
- habitatData — S_HabitatData structure array that stores data for each habitat in the population.
- probabilities — array of probabilities used for mutation.
Private methods contain the implementation of the main steps of the BBO algorithm, such as population initialization, calculation of migration rates and mutation.
//—————————————————————————————————————————————————————————————————————————————— class C_AO_BBO : public C_AO { public: //-------------------------------------------------------------------- ~C_AO_BBO () { } C_AO_BBO () { ao_name = "BBO"; ao_desc = "Biogeography-Based Optimization"; ao_link = "https://www.mql5.com/en/articles/18354"; popSize = 50; // population size (number of habitats) immigrationMax = 1.0; // maximum immigration rate emigrationMax = 1.0; // maximum emigration rate mutationProb = 0.5; // mutation probability elitismCount = 2; // number of elite solutions speciesMax = 50; // maximum number of species ArrayResize (params, 6); params [0].name = "popSize"; params [0].val = popSize; params [1].name = "immigrationMax"; params [1].val = immigrationMax; params [2].name = "emigrationMax"; params [2].val = emigrationMax; params [3].name = "mutationProb"; params [3].val = mutationProb; params [4].name = "elitismCount"; params [4].val = elitismCount; params [5].name = "speciesMax"; params [5].val = speciesMax; } void SetParams () { popSize = (int)params [0].val; immigrationMax = params [1].val; emigrationMax = params [2].val; mutationProb = params [3].val; elitismCount = (int)params [4].val; speciesMax = (int)params [5].val; } bool Init (const double &rangeMinP [], // minimum values const double &rangeMaxP [], // maximum values const double &rangeStepP [], // step change const int epochsP = 0); // number of epochs void Moving (); void Revision (); //---------------------------------------------------------------------------- double immigrationMax; // maximum immigration rate double emigrationMax; // maximum emigration rate double mutationProb; // mutation probability int elitismCount; // number of elite solutions int speciesMax; // maximum number of species private: //------------------------------------------------------------------- struct S_HabitatData { int speciesCount; // number of species in the habitat double immigration; // immigration rate double emigration; // emigration rate double probability; // probability of existence }; S_HabitatData habitatData []; // data for each habitat double probabilities []; // probabilities for counting mutations // Auxiliary methods void InitializePopulation (); void CalculateRates (); void Migration (); void Mutation (); double CalculateProbability (int speciesCount); }; //——————————————————————————————————————————————————————————————————————————————
The Init method configures the BBO algorithm before it starts working. It performs basic initialization (checks and settings), allocates memory to store data about "habitats" and migration probabilities. It then calculates and normalizes migration probabilities based on the number of species in each habitat. Returns 'true' on success.
//—————————————————————————————————————————————————————————————————————————————— bool C_AO_BBO::Init (const double &rangeMinP [], // minimum values const double &rangeMaxP [], // maximum values const double &rangeStepP [], // step change const int epochsP = 0) // number of epochs { if (!StandardInit (rangeMinP, rangeMaxP, rangeStepP)) return false; //---------------------------------------------------------------------------- // Initialize arrays for each habitat ArrayResize (habitatData, popSize); ArrayResize (probabilities, speciesMax + 1); // Calculate probabilities for different numbers of species double sum = 0.0; for (int i = 0; i <= speciesMax; i++) { probabilities [i] = CalculateProbability (i); sum += probabilities [i]; } // Normalization of probabilities if (sum > 0) { for (int i = 0; i <= speciesMax; i++) { probabilities [i] /= sum; } } return true; } //——————————————————————————————————————————————————————————————————————————————
The Moving method implements the main optimization cycle of the BBO algorithm. The first time the method is called, the "revision" flag is checked. If it is set to a value indicating that the population has not yet been created, it is initialized. This involves generating random solutions, assessing their suitability, and setting initial parameters. After this, the flag is reset.
Once the initialization is complete, a series of steps characteristic of an algorithmic cycle are performed: the population of solutions is sorted by their fitness value to identify the best and worst agents. This helps manage migrations and mutations. At this stage, the probabilities and rates of species exchange between habitats are calculated based on their current state and the quality of their decisions. These parameters control how species "migrate" from one habitat to another. In this step, the exchange of SIV between habitats occurs.
As a result, diversity-poor sites receive new species from richer options, facilitating exploration of the solution space. After migration, random changes in solutions (mutation) occur to maintain genetic diversity. Mutation probabilities may depend on the current state of the solution and the algorithm parameters. At the end of the loop, the current values of the solution fitness function are saved so that they can be used for sorting and analysis in the next iteration of the algorithm.
//+----------------------------------------------------------------------------+ //| Basic optimization method | //+----------------------------------------------------------------------------+ void C_AO_BBO::Moving () { // First iteration - initialization of the initial population if (!revision) { InitializePopulation (); revision = true; return; } // Main optimization // 1. Sort the population by HSI (fitness) static S_AO_Agent aTemp []; ArrayResize (aTemp, popSize); u.Sorting (a, aTemp, popSize); // 2. Calculate immigration and emigration rates CalculateRates (); // 3. Migration (exchange of SIVs between habitats) Migration (); // 4. Probability-based mutation Mutation (); // 5. Save state for the next iteration for (int i = 0; i < popSize; i++) { a [i].fP = a [i].f; } } //——————————————————————————————————————————————————————————————————————————————
The Revision method is designed to update the current best solution in the population. It goes through all agents (solutions) of the current population and compares their fitness function value with the one stored in the fB variable, which stores the best result at the moment.
If any agent has a fitness function value better than the current best one, it replaces that value, and the corresponding solution parameters are copied into the variable storing the best solution. As a result, after executing the method, the variable always contains the best solution found.
//+----------------------------------------------------------------------------+ //| Update the best solution | //+----------------------------------------------------------------------------+ void C_AO_BBO::Revision () { // Find the best solution in the current population for (int i = 0; i < popSize; i++) { // Update the best solution if (a [i].f > fB) { fB = a [i].f; ArrayCopy (cB, a [i].c, 0, 0, WHOLE_ARRAY); } } } //——————————————————————————————————————————————————————————————————————————————
The InitializePopulation method is responsible for creating the initial population of solutions for the BBO algorithm. It creates popSize (population size) of individuals (habitats) uniformly distributed throughout the search space.
For each individual, the method generates random coordinates (values of the solution parameters) within the limits specified by the rangeMin (minimum boundaries) and rangeMax (maximum boundaries) arrays for each "coords" coordinate. The u.RNDfromCI function is used to generate a random number in a given range.
Next, the method rounds the generated coordinates to the nearest valid step defined by the rangeStep array. This ensures that the solutions are in a feasible discrete search space. The SeInDiSp function is used for this. After initializing the coordinates, the method initializes the habitatData data structure for each individual, setting the speciesCount, immigration, emigration and probability values to zero. These values will be used in the optimization to calculate immigration, emigration rates, and mutation probabilities.
//+----------------------------------------------------------------------------+ //| Initialize the initial population | //+----------------------------------------------------------------------------+ void C_AO_BBO::InitializePopulation () { // Initialize the initial population uniformly throughout the space for (int i = 0; i < popSize; i++) { for (int c = 0; c < coords; c++) { // Generate random coordinates within acceptable limits a [i].c [c] = u.RNDfromCI (rangeMin [c], rangeMax [c]); // Round to the nearest acceptable step a [i].c [c] = u.SeInDiSp (a [i].c [c], rangeMin [c], rangeMax [c], rangeStep [c]); } // Initialize habitat data habitatData [i].speciesCount = 0; habitatData [i].immigration = 0.0; habitatData [i].emigration = 0.0; habitatData [i].probability = 0.0; } } //——————————————————————————————————————————————————————————————————————————————
The CalculateRates method is designed to calculate migration rates (immigration and emigration) and the probability of existence of each habitat in the population.
A linear model is used, in which the number of species associated with each solution is determined proportionally to its rank, with better solutions having more species. The immigration rate decreases with increasing number of species — the more species a solution has, the lower its probability of accepting new individuals. The rate of emigration increases with the number of species - the more species a solution has, the higher the probability of leaving it.
The probability of habitat existence is established based on predetermined probabilities for each number of species. If the number of species exceeds the maximum allowed, the probability is set to zero.
//+----------------------------------------------------------------------------+ //| Calculate immigration and emigration rates | //+----------------------------------------------------------------------------+ void C_AO_BBO::CalculateRates () { // For the linear migration model for (int i = 0; i < popSize; i++) { // The number of species is inversely proportional to the rank (the best solutions have more species) habitatData [i].speciesCount = speciesMax - (i * speciesMax / popSize); // The rate of immigration decreases as the number of species increases habitatData [i].immigration = immigrationMax * (1.0 - (double)habitatData [i].speciesCount / speciesMax); // The rate of emigration increases with the number of species habitatData [i].emigration = emigrationMax * (double)habitatData [i].speciesCount / speciesMax; // Probability of habitat existence if (habitatData [i].speciesCount <= speciesMax) { habitatData [i].probability = probabilities [habitatData [i].speciesCount]; } else { habitatData [i].probability = 0.0; } } } //——————————————————————————————————————————————————————————————————————————————
The Migration method implements the process of migration (exchange of SIV - Habitat Suitability Index Variables, i.e. coordinate values) between habitats in a population. The idea behind the method is that habitats with high immigration rates (i.e. those with few species) can "accept" SIVs from other habitats with high emigration rates (those with many species).
The loop iterates through all habitats in the population, but skips the first elitismCount habitats, which are considered "elite" and not subject to migration. For each habitat (except elite ones), it is randomly determined whether it will be modified in the current iteration. The probability of modification is determined by the value of "habitatData[i].immigration" (the immigration rate of the current habitat). If a habitat is selected for migration, the algorithm iterates over all of its coordinates (SIV). For each coordinate, it is again randomly determined whether it will be modified. The probability of modification is also determined by the value of "habitatData[i].immigration".
If a coordinate is selected for modification, the habitat the value of this coordinate will be "borrowed" from is determined. This selection is based on roulette wheel selection proportional to the values of "habitatData[j].emigration" (emigration rates of other habitats). That is, habitats with higher emigration rates have a greater chance of becoming a source of migration. When calculating probabilities, it is taken into account that the current habitat is not chosen as a source for itself. If a migration source is selected, the value of the corresponding coordinate (SIV) is copied from the source habitat to the current habitat.
Ultimately, the method performs a regulated information exchange (SIV) process between habitats, where habitats with low species abundance (high immigration) receive information from habitats with high species abundance (high emigration). This allows good (SIV) solutions to be spread throughout the population while keeping the best solutions unchanged.
//+----------------------------------------------------------------------------+ //| Migration (exchange of SIVs between habitats) | //+----------------------------------------------------------------------------+ void C_AO_BBO::Migration () { for (int i = 0; i < popSize; i++) { // Skip elite solutions if (i < elitismCount) continue; // Determine whether the habitat will be modified if (u.RNDprobab () < habitatData [i].immigration) { // For each coordinate (SIV) for (int c = 0; c < coords; c++) { // Determine whether this coordinate will be modified if (u.RNDprobab () < habitatData [i].immigration) { // Select a migration source based on emigration rates double sumEmigration = 0.0; for (int j = 0; j < popSize; j++) { if (j != i) sumEmigration += habitatData [j].emigration; } if (sumEmigration > 0) { // Roulette source selection double roulette = u.RNDprobab () * sumEmigration; double cumSum = 0.0; for (int j = 0; j < popSize; j++) { if (j != i) { cumSum += habitatData [j].emigration; if (roulette <= cumSum) { // Copy SIV from habitat j to habitat i a [i].c [c] = a [j].c [c]; break; } } } } } } } } } //——————————————————————————————————————————————————————————————————————————————
The Mutation method mutates habitats in a population. The goal of mutation is to introduce random changes to solutions to avoid getting stuck in local optima and explore new search space.
As in the migration method, elite solutions (the first elitismCount habitats) are skipped and not mutated. This is necessary to save the best solutions found. For each remaining habitat, the mutation probability is calculated. It is important that mutation probability is inversely proportional to the habitat existence probability (habitatData [i].probability). This means that habitats with low existence probability — including both very poor and very strong extreme solutions — mutate more often to facilitate exploration of new areas.
If the generated random number is less than the calculated mutationRate, then a mutation occurs. One mutateCoord (SIV) coordinate is randomly selected for mutation from among all "coords". A new random value within the specified range is generated for the selected coordinate. Next, the SeInDiSp function is applied to the new value, limiting the value to the specified limits.
Thus, the Mutation method introduces an element of randomness into the search process, allowing the algorithm to find new, potentially better solutions, especially in areas that have not yet been sufficiently explored. The mutation rate is adjusted by the probability of habitat existence to balance exploration and exploitation (exploration vs. exploitation).
//+----------------------------------------------------------------------------+ //| Probability-based mutation | //+----------------------------------------------------------------------------+ void C_AO_BBO::Mutation () { for (int i = 0; i < popSize; i++) { // Skip elite solutions if (i < elitismCount) continue; // The mutation rate is inversely proportional to the probability of existence double mutationRate = mutationProb * (1.0 - habitatData [i].probability); if (u.RNDprobab () < mutationRate) { // Select a random coordinate for mutation int mutateCoord = MathRand () % coords; // Generate a new value for the selected coordinate a [i].c [mutateCoord] = u.RNDfromCI (rangeMin [mutateCoord], rangeMax [mutateCoord]); a [i].c [mutateCoord] = u.SeInDiSp (a [i].c [mutateCoord], rangeMin [mutateCoord], rangeMax [mutateCoord], rangeStep [mutateCoord]); } } } //——————————————————————————————————————————————————————————————————————————————
The CalculateProbability method calculates the probability of a habitat existing given the number of species. The method uses a simplified model: the maximum probability is achieved near the equilibrium value of the species (in the middle of the range), and when deviating from it, the probability rapidly decreases along a Gaussian curve. The further speciesCount is from speciesMax/2, the lower the probability.
Overall, the method produces a model, in which habitats with species numbers close to the equilibrium value have a higher probability of existing than habitats with species numbers that deviate greatly from the equilibrium value. This represents a simplified but effective model of habitat "suitability" based on its biological diversity.
//+----------------------------------------------------------------------------+ //| Calculate the probability for a given number of species | //+----------------------------------------------------------------------------+ double C_AO_BBO::CalculateProbability (int speciesCount) { // Simplified probability model // Maximum probability in the middle of the range (equilibrium) int equilibrium = speciesMax / 2; double distance = MathAbs (speciesCount - equilibrium); double probability = MathExp (-distance * distance / (2.0 * equilibrium * equilibrium)); return probability; } //——————————————————————————————————————————————————————————————————————————————
Test results
After experimenting a bit with the parameters, I have discovered excellent results from the BBO algorithm.
BBO|Biogeography-Based Optimization|50.0|1.0|1.0|0.5|2.0|50.0|
=============================
5 Hilly's; Func runs: 10000; result: 0.9491244808033844
25 Hilly's; Func runs: 10000; result: 0.6945610309062928
500 Hilly's; Func runs: 10000; result: 0.35031241665471596
=============================
5 Forest's; Func runs: 10000; result: 0.9381951766964413
25 Forest's; Func runs: 10000; result: 0.6736501622157315
500 Forest's; Func runs: 10000; result: 0.2568167323109364
=============================
5 Megacity's; Func runs: 10000; result: 0.7461538461538464
25 Megacity's; Func runs: 10000; result: 0.4827692307692309
500 Megacity's; Func runs: 10000; result: 0.17369230769230892
=============================
All score: 5.26528 (58.50%)
The visualization shows how efficient the BBO algorithm is. On the most difficult Megacity discrete function, the algorithm shows excellent results.
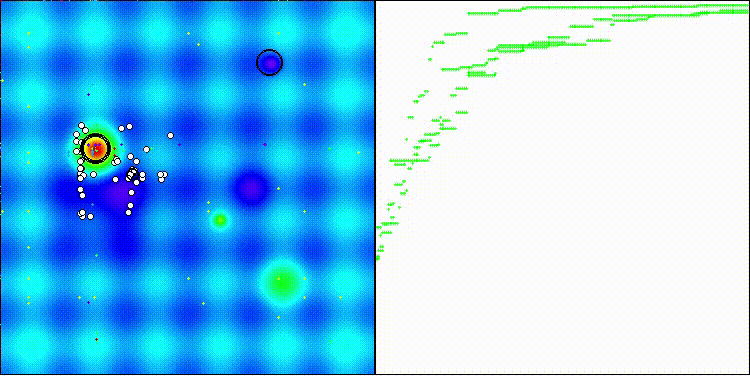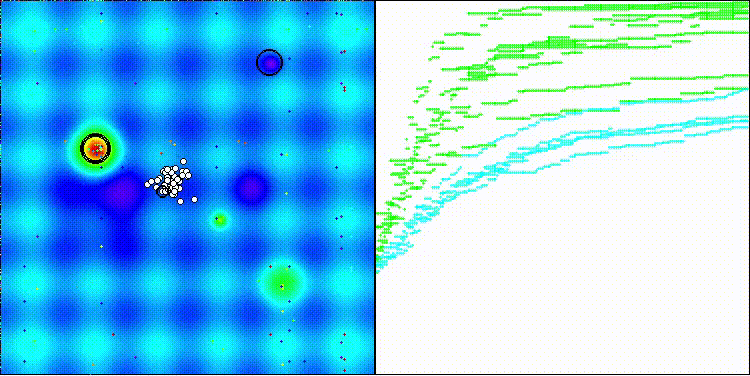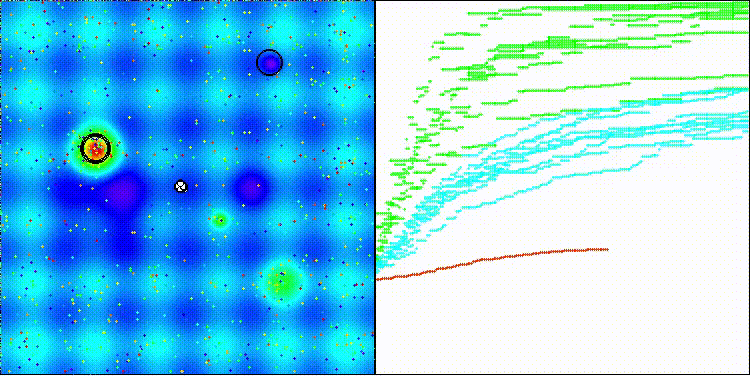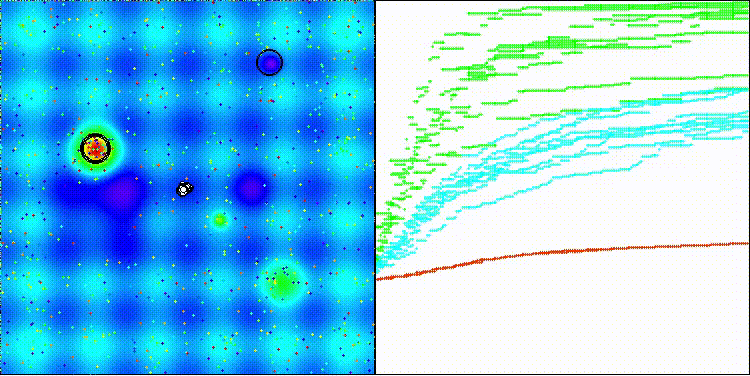
BBO on the Hilly test function
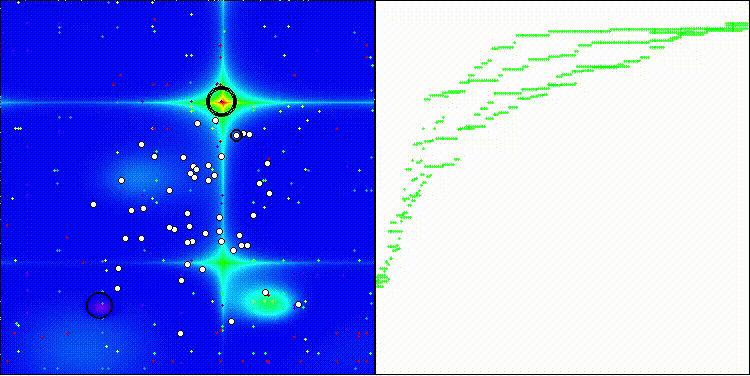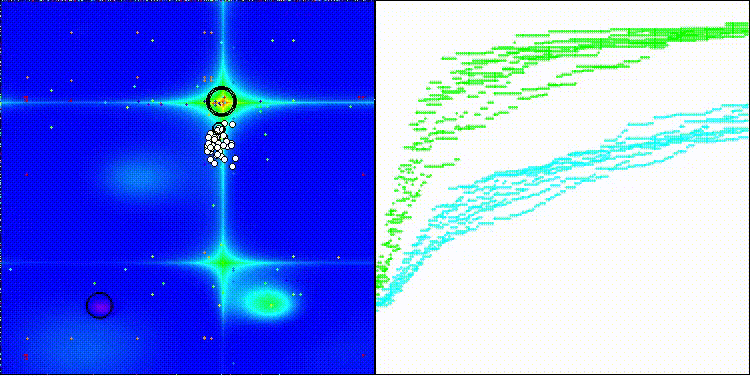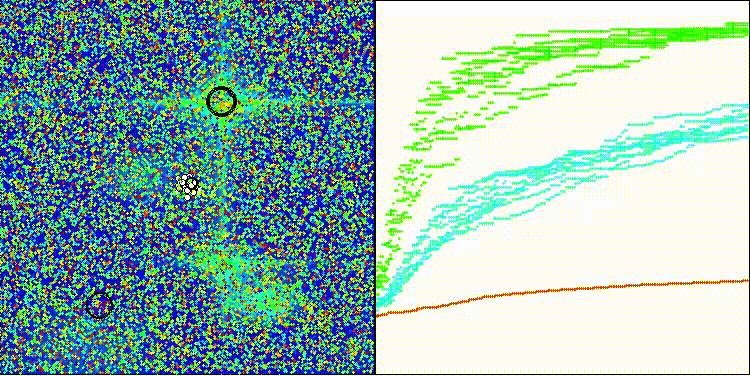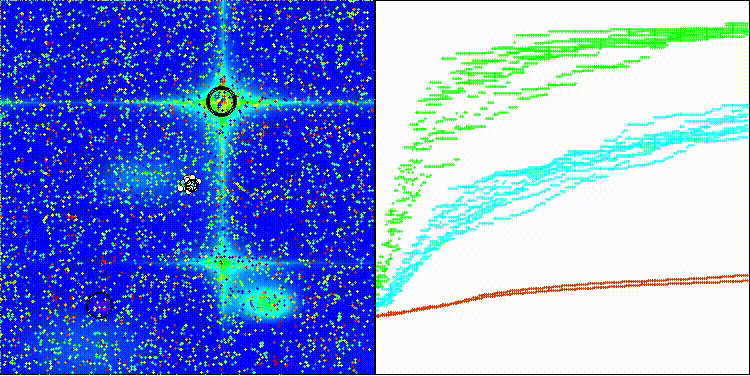
BBO on the Forest test function

BBO on the Megacity test function
Based on the test results, the elegant BBO algorithm occupies the high 12th place at the top of the rating table.
| # | AO | Description | Hilly | Hilly Final | Forest | Forest Final | Megacity (discrete) | Megacity Final | Final Result | % of MAX | ||||||
| 10 p (5 F) | 50 p (25 F) | 1000 p (500 F) | 10 p (5 F) | 50 p (25 F) | 1000 p (500 F) | 10 p (5 F) | 50 p (25 F) | 1000 p (500 F) | ||||||||
| 1 | ANS | across neighbourhood search | 0.94948 | 0.84776 | 0.43857 | 2.23581 | 1.00000 | 0.92334 | 0.39988 | 2.32323 | 0.70923 | 0.63477 | 0.23091 | 1.57491 | 6.134 | 68.15 |
| 2 | CLA | code lock algorithm (joo) | 0.95345 | 0.87107 | 0.37590 | 2.20042 | 0.98942 | 0.91709 | 0.31642 | 2.22294 | 0.79692 | 0.69385 | 0.19303 | 1.68380 | 6.107 | 67.86 |
| 3 | AMOm | animal migration ptimization M | 0.90358 | 0.84317 | 0.46284 | 2.20959 | 0.99001 | 0.92436 | 0.46598 | 2.38034 | 0.56769 | 0.59132 | 0.23773 | 1.39675 | 5.987 | 66.52 |
| 4 | (P+O)ES | (P+O) evolution strategies | 0.92256 | 0.88101 | 0.40021 | 2.20379 | 0.97750 | 0.87490 | 0.31945 | 2.17185 | 0.67385 | 0.62985 | 0.18634 | 1.49003 | 5.866 | 65.17 |
| 5 | CTA | comet tail algorithm (joo) | 0.95346 | 0.86319 | 0.27770 | 2.09435 | 0.99794 | 0.85740 | 0.33949 | 2.19484 | 0.88769 | 0.56431 | 0.10512 | 1.55712 | 5.846 | 64.96 |
| 6 | TETA | time evolution travel algorithm (joo) | 0.91362 | 0.82349 | 0.31990 | 2.05701 | 0.97096 | 0.89532 | 0.29324 | 2.15952 | 0.73462 | 0.68569 | 0.16021 | 1.58052 | 5.797 | 64.41 |
| 7 | SDSm | stochastic diffusion search M | 0.93066 | 0.85445 | 0.39476 | 2.17988 | 0.99983 | 0.89244 | 0.19619 | 2.08846 | 0.72333 | 0.61100 | 0.10670 | 1.44103 | 5.709 | 63.44 |
| 8 | BOAm | billiards optimization algorithm M | 0.95757 | 0.82599 | 0.25235 | 2.03590 | 1.00000 | 0.90036 | 0.30502 | 2.20538 | 0.73538 | 0.52523 | 0.09563 | 1.35625 | 5.598 | 62.19 |
| 9 | AAm | archery algorithm M | 0.91744 | 0.70876 | 0.42160 | 2.04780 | 0.92527 | 0.75802 | 0.35328 | 2.03657 | 0.67385 | 0.55200 | 0.23738 | 1.46323 | 5.548 | 61.64 |
| 10 | ESG | evolution of social groups (joo) | 0.99906 | 0.79654 | 0.35056 | 2.14616 | 1.00000 | 0.82863 | 0.13102 | 1.95965 | 0.82333 | 0.55300 | 0.04725 | 1.42358 | 5.529 | 61.44 |
| 11 | SIA | simulated isotropic annealing (joo) | 0.95784 | 0.84264 | 0.41465 | 2.21513 | 0.98239 | 0.79586 | 0.20507 | 1.98332 | 0.68667 | 0.49300 | 0.09053 | 1.27020 | 5.469 | 60.76 |
| 12 | BBO | biogeography based optimization | 0.94912 | 0.69456 | 0.35031 | 1.99399 | 0.93820 | 0.67365 | 0.25682 | 1.86867 | 0.74615 | 0.48277 | 0.17369 | 1.40261 | 5.265 | 58.50 |
| 13 | ACS | artificial cooperative search | 0.75547 | 0.74744 | 0.30407 | 1.80698 | 1.00000 | 0.88861 | 0.22413 | 2.11274 | 0.69077 | 0.48185 | 0.13322 | 1.30583 | 5.226 | 58.06 |
| 14 | DA | dialectical algorithm | 0.86183 | 0.70033 | 0.33724 | 1.89940 | 0.98163 | 0.72772 | 0.28718 | 1.99653 | 0.70308 | 0.45292 | 0.16367 | 1.31967 | 5.216 | 57.95 |
| 15 | BHAm | black hole algorithm M | 0.75236 | 0.76675 | 0.34583 | 1.86493 | 0.93593 | 0.80152 | 0.27177 | 2.00923 | 0.65077 | 0.51646 | 0.15472 | 1.32195 | 5.196 | 57.73 |
| 16 | ASO | anarchy society optimization | 0.84872 | 0.74646 | 0.31465 | 1.90983 | 0.96148 | 0.79150 | 0.23803 | 1.99101 | 0.57077 | 0.54062 | 0.16614 | 1.27752 | 5.178 | 57.54 |
| 17 | RFO | royal flush optimization (joo) | 0.83361 | 0.73742 | 0.34629 | 1.91733 | 0.89424 | 0.73824 | 0.24098 | 1.87346 | 0.63154 | 0.50292 | 0.16421 | 1.29867 | 5.089 | 56.55 |
| 18 | AOSm | atomic orbital search M | 0.80232 | 0.70449 | 0.31021 | 1.81702 | 0.85660 | 0.69451 | 0.21996 | 1.77107 | 0.74615 | 0.52862 | 0.14358 | 1.41835 | 5.006 | 55.63 |
| 19 | TSEA | turtle shell evolution algorithm (joo) | 0.96798 | 0.64480 | 0.29672 | 1.90949 | 0.99449 | 0.61981 | 0.22708 | 1.84139 | 0.69077 | 0.42646 | 0.13598 | 1.25322 | 5.004 | 55.60 |
| 20 | DE | differential evolution | 0.95044 | 0.61674 | 0.30308 | 1.87026 | 0.95317 | 0.78896 | 0.16652 | 1.90865 | 0.78667 | 0.36033 | 0.02953 | 1.17653 | 4.955 | 55.06 |
| 21 | SRA | successful restaurateur algorithm (joo) | 0.96883 | 0.63455 | 0.29217 | 1.89555 | 0.94637 | 0.55506 | 0.19124 | 1.69267 | 0.74923 | 0.44031 | 0.12526 | 1.31480 | 4.903 | 54.48 |
| 22 | CRO | chemical reaction optimization | 0.94629 | 0.66112 | 0.29853 | 1.90593 | 0.87906 | 0.58422 | 0.21146 | 1.67473 | 0.75846 | 0.42646 | 0.12686 | 1.31178 | 4.892 | 54.36 |
| 23 | BIO | blood inheritance optimization (joo) | 0.81568 | 0.65336 | 0.30877 | 1.77781 | 0.89937 | 0.65319 | 0.21760 | 1.77016 | 0.67846 | 0.47631 | 0.13902 | 1.29378 | 4.842 | 53.80 |
| 24 | BSA | bird swarm algorithm | 0.89306 | 0.64900 | 0.26250 | 1.80455 | 0.92420 | 0.71121 | 0.24939 | 1.88479 | 0.69385 | 0.32615 | 0.10012 | 1.12012 | 4.809 | 53.44 |
| 25 | HS | harmony search | 0.86509 | 0.68782 | 0.32527 | 1.87818 | 0.99999 | 0.68002 | 0.09590 | 1.77592 | 0.62000 | 0.42267 | 0.05458 | 1.09725 | 4.751 | 52.79 |
| 26 | SSG | saplings sowing and growing | 0.77839 | 0.64925 | 0.39543 | 1.82308 | 0.85973 | 0.62467 | 0.17429 | 1.65869 | 0.64667 | 0.44133 | 0.10598 | 1.19398 | 4.676 | 51.95 |
| 27 | BCOm | bacterial chemotaxis optimization M | 0.75953 | 0.62268 | 0.31483 | 1.69704 | 0.89378 | 0.61339 | 0.22542 | 1.73259 | 0.65385 | 0.42092 | 0.14435 | 1.21912 | 4.649 | 51.65 |
| 28 | ABO | african buffalo optimization | 0.83337 | 0.62247 | 0.29964 | 1.75548 | 0.92170 | 0.58618 | 0.19723 | 1.70511 | 0.61000 | 0.43154 | 0.13225 | 1.17378 | 4.634 | 51.49 |
| 29 | (PO)ES | (PO) evolution strategies | 0.79025 | 0.62647 | 0.42935 | 1.84606 | 0.87616 | 0.60943 | 0.19591 | 1.68151 | 0.59000 | 0.37933 | 0.11322 | 1.08255 | 4.610 | 51.22 |
| 30 | FBA | Fractal-Based Algorithm | 0.79000 | 0.65134 | 0.28965 | 1.73099 | 0.87158 | 0.56823 | 0.18877 | 1.62858 | 0.61077 | 0.46062 | 0.12398 | 1.19537 | 4.555 | 50.61 |
| 31 | TSm | tabu search M | 0.87795 | 0.61431 | 0.29104 | 1.78330 | 0.92885 | 0.51844 | 0.19054 | 1.63783 | 0.61077 | 0.38215 | 0.12157 | 1.11449 | 4.536 | 50.40 |
| 32 | BSO | brain storm optimization | 0.93736 | 0.57616 | 0.29688 | 1.81041 | 0.93131 | 0.55866 | 0.23537 | 1.72534 | 0.55231 | 0.29077 | 0.11914 | 0.96222 | 4.498 | 49.98 |
| 33 | WOAm | wale optimization algorithm M | 0.84521 | 0.56298 | 0.26263 | 1.67081 | 0.93100 | 0.52278 | 0.16365 | 1.61743 | 0.66308 | 0.41138 | 0.11357 | 1.18803 | 4.476 | 49.74 |
| 34 | AEFA | artificial electric field algorithm | 0.87700 | 0.61753 | 0.25235 | 1.74688 | 0.92729 | 0.72698 | 0.18064 | 1.83490 | 0.66615 | 0.11631 | 0.09508 | 0.87754 | 4.459 | 49.55 |
| 35 | AEO | artificial ecosystem-based optimization algorithm | 0.91380 | 0.46713 | 0.26470 | 1.64563 | 0.90223 | 0.43705 | 0.21400 | 1.55327 | 0.66154 | 0.30800 | 0.28563 | 1.25517 | 4.454 | 49.49 |
| 36 | CAm | camel algorithm M | 0.78684 | 0.56042 | 0.35133 | 1.69859 | 0.82772 | 0.56041 | 0.24336 | 1.63149 | 0.64846 | 0.33092 | 0.13418 | 1.11356 | 4.444 | 49.37 |
| 37 | ACOm | ant colony optimization M | 0.88190 | 0.66127 | 0.30377 | 1.84693 | 0.85873 | 0.58680 | 0.15051 | 1.59604 | 0.59667 | 0.37333 | 0.02472 | 0.99472 | 4.438 | 49.31 |
| 38 | BFO-GA | bacterial foraging optimization - ga | 0.89150 | 0.55111 | 0.31529 | 1.75790 | 0.96982 | 0.39612 | 0.06305 | 1.42899 | 0.72667 | 0.27500 | 0.03525 | 1.03692 | 4.224 | 46.93 |
| 39 | SOA | simple optimization algorithm | 0.91520 | 0.46976 | 0.27089 | 1.65585 | 0.89675 | 0.37401 | 0.16984 | 1.44060 | 0.69538 | 0.28031 | 0.10852 | 1.08422 | 4.181 | 46.45 |
| 40 | ABHA | artificial bee hive algorithm | 0.84131 | 0.54227 | 0.26304 | 1.64663 | 0.87858 | 0.47779 | 0.17181 | 1.52818 | 0.50923 | 0.33877 | 0.10397 | 0.95197 | 4.127 | 45.85 |
| 41 | ACMO | atmospheric cloud model optimization | 0.90321 | 0.48546 | 0.30403 | 1.69270 | 0.80268 | 0.37857 | 0.19178 | 1.37303 | 0.62308 | 0.24400 | 0.10795 | 0.97503 | 4.041 | 44.90 |
| 42 | ADAMm | adaptive moment estimation M | 0.88635 | 0.44766 | 0.26613 | 1.60014 | 0.84497 | 0.38493 | 0.16889 | 1.39880 | 0.66154 | 0.27046 | 0.10594 | 1.03794 | 4.037 | 44.85 |
| 43 | CGO | chaos game optimization | 0.57256 | 0.37158 | 0.32018 | 1.26432 | 0.61176 | 0.61931 | 0.62161 | 1.85267 | 0.37538 | 0.21923 | 0.19028 | 0.78490 | 3.902 | 43.35 |
| 44 | CROm | coral reefs optimization M | 0.78512 | 0.46032 | 0.25958 | 1.50502 | 0.86688 | 0.35297 | 0.16267 | 1.38252 | 0.63231 | 0.26738 | 0.10734 | 1.00703 | 3.895 | 43.27 |
| 45 | ATAm | artificial tribe algorithm M | 0.71771 | 0.55304 | 0.25235 | 1.52310 | 0.82491 | 0.55904 | 0.20473 | 1.58867 | 0.44000 | 0.18615 | 0.09411 | 0.72026 | 3.832 | 42.58 |
| RW | neuroboids optimization algorithm 2(joo) | 0.48754 | 0.32159 | 0.25781 | 1.06694 | 0.37554 | 0.21944 | 0.15877 | 0.75375 | 0.27969 | 0.14917 | 0.09847 | 0.52734 | 2.348 | 26.09 | |
Summary
The Biogeography-Based Optimization (BBO) algorithm demonstrated impressive results in the tests, ranking 12th among the top 45 population-based optimization algorithms with an overall performance score of 58.5%. This is an exceptional result for an algorithm based on such an elegant and intuitive natural metaphor.
Particularly noteworthy is BBO's ability to handle medium- and high-dimensional problems efficiently, demonstrating its scalability and resistance to the "curse of dimensionality" — a problem faced by many optimization algorithms.
The conceptual basis of BBO — the migration of species between islands — proved to be not just a beautiful metaphor, but also a highly efficient mechanism for balancing the exploration of the search space and the exploitation of the solutions found. A linear migration model, where rich habitats actively share their characteristics and poor habitats actively adopt them, creates a natural gradient of information flow from better solutions to worse ones.
The BBO mutation operator deserves special attention, as it is fundamentally different from classical approaches. Instead of a fixed or random probability of mutation, BBO uses a theoretically based model where the probability of mutation is inversely proportional to the probability of habitat existence. This means that the most "natural" and stable solutions (with an average number of species) mutate rarely, while extreme solutions - both very good and very bad - undergo more frequent changes. This approach creates an adaptive mechanism that automatically regulates the balance between stability and variability of the population.
The algorithm demonstrates excellent stability across various landscape types: it is uniformly green in color across all tests and shows no significant drop in results compared to other optimization algorithms.
An important advantage of BBO is its conceptual simplicity coupled with high efficiency. Unlike some modern metaheuristics that require many parameters and complex operators, BBO operates with intuitive concepts: migration, number of species, habitat suitability.
The test results confirm that BBO is not just "yet another" bioinspired algorithm, but a fully-fledged and competitive optimization method capable of competing with recognized leaders in the field. The combination of theoretical soundness, computational efficiency, and practical applicability makes BBO a valuable addition to the arsenal of modern global optimization methods.
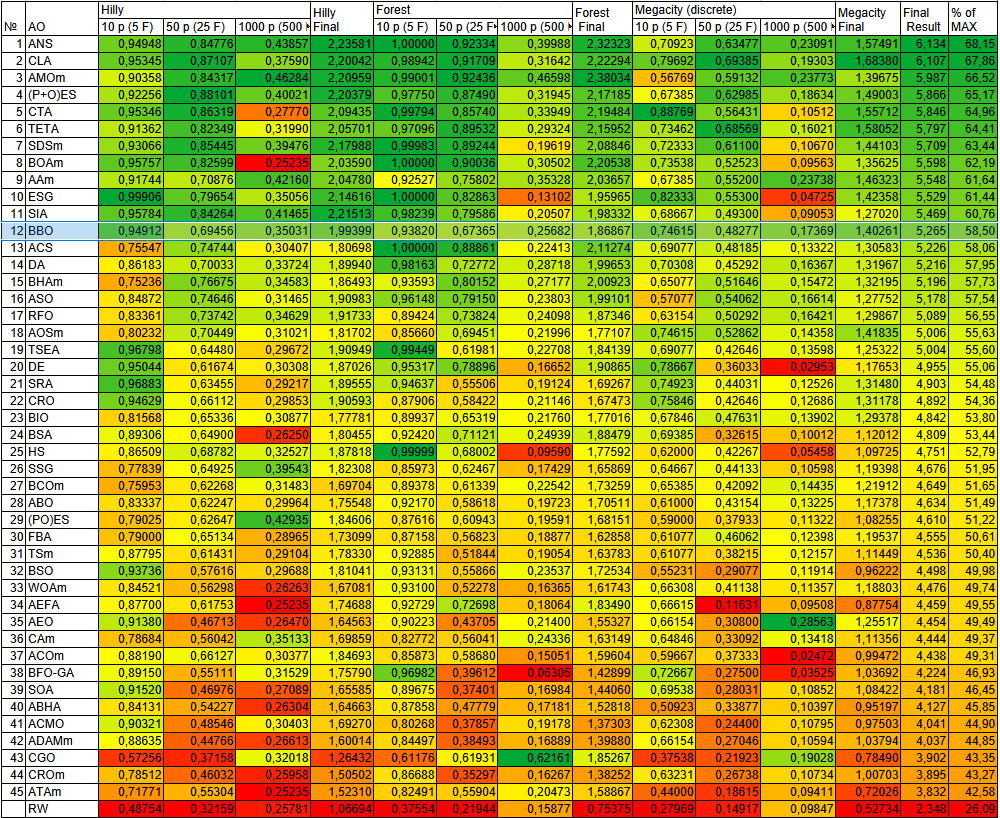
Figure 2. Color gradation of algorithms according to the corresponding tests
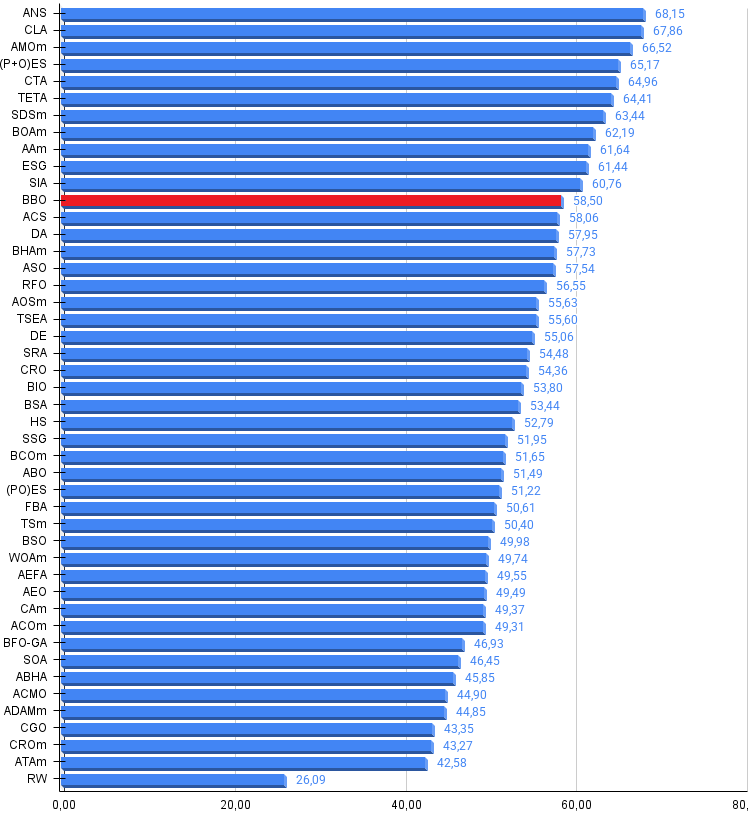
Figure 3. Histogram of algorithm testing results (scale from 0 to 100, the higher the better, where 100 is the maximum possible theoretical result, in the archive there is a script for calculating the rating table)
BBO pros and cons:
Pros:
- Fast.
- Simple implementation.
- Good results across a wide range of problem dimensions.
Cons:
- A large number of parameters.
The article is accompanied by an archive with the current versions of the algorithm codes. The author of the article is not responsible for the absolute accuracy in the description of canonical algorithms. Changes have been made to many of them to improve search capabilities. The conclusions and judgments presented in the articles are based on the results of the experiments.
Programs used in the article
| # | Name | Type | Description |
|---|---|---|---|
| 1 | #C_AO.mqh | Include | Parent class of population optimization algorithms |
| 2 | #C_AO_enum.mqh | Include | Enumeration of population optimization algorithms |
| 3 | TestFunctions.mqh | Include | Library of test functions |
| 4 | TestStandFunctions.mqh | Include | Test stand function library |
| 5 | Utilities.mqh | Include | Library of auxiliary functions |
| 6 | CalculationTestResults.mqh | Include | Script for calculating results in the comparison table |
| 7 | Testing AOs.mq5 | Script | The unified test stand for all population optimization algorithms |
| 8 | Simple use of population optimization algorithms.mq5 | Script | A simple example of using population optimization algorithms without visualization |
| 9 | Test_AO_BBO.mq5 | Script | BBO test stand |
Translated from Russian by MetaQuotes Ltd.
Original article: https://www.mql5.com/ru/articles/18354
Warning: All rights to these materials are reserved by MetaQuotes Ltd. Copying or reprinting of these materials in whole or in part is prohibited.
This article was written by a user of the site and reflects their personal views. MetaQuotes Ltd is not responsible for the accuracy of the information presented, nor for any consequences resulting from the use of the solutions, strategies or recommendations described.
 Manual Backtesting with On-Chart Buttons in the MetaTrader 5 Strategy Tester
Manual Backtesting with On-Chart Buttons in the MetaTrader 5 Strategy Tester
 Leak-Free Multi-Timeframe Engine with Closed-Bar Reads in MQL5
Leak-Free Multi-Timeframe Engine with Closed-Bar Reads in MQL5
 Building a Trade Analytics System (Part 3): Storing MetaTrader 5 Trades in SQLite
Building a Trade Analytics System (Part 3): Storing MetaTrader 5 Trades in SQLite
 From Matrices to Models: How to Build an ML Pipeline in MQL5 and Export It to ONNX
From Matrices to Models: How to Build an ML Pipeline in MQL5 and Export It to ONNX
- Free trading apps
- Over 8,000 signals for copying
- Economic news for exploring financial markets
You agree to website policy and terms of use