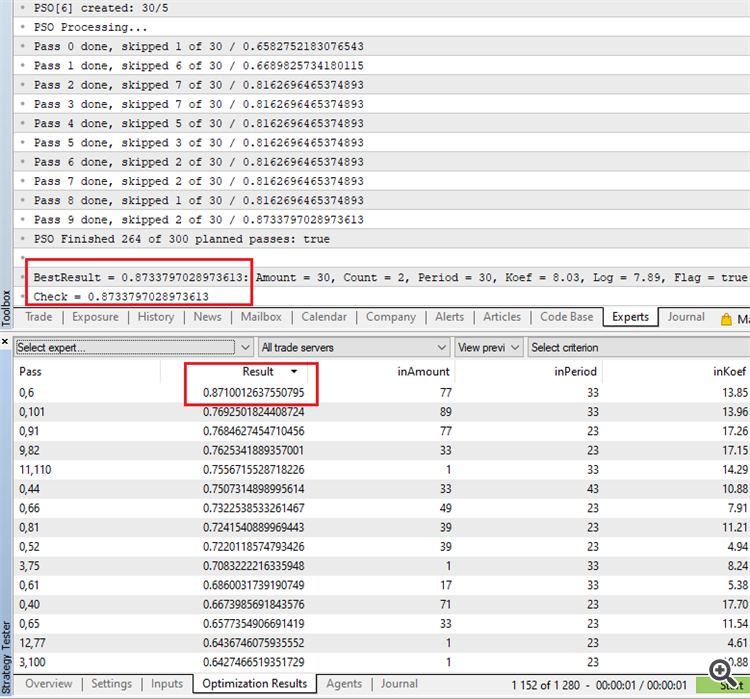
MQL's OOP notes: Particle Swarm Speeds Up Expert Adviser Optimization On The Fly
There is a number of methods of fast optimization. One of them is genetic optimization, which is used by the tester in MetaTrader. The other one is simulated annealing. And yet another one is particle swarm optimization. This method is relatively simple to implement. Let's code it in MQL.

According to the algorithm every particle should have current position, velocity, and memory about its best position in past. The "best" means that EA demonstarted highest (known to the particle) performance at those position (parameter set). Let's put this in a class.
{
public:
double position[]; // current point
double best[]; // best point known to the particle
double velocity[]; // current speed
double positionValue; // EA performance in current point
double bestValue; // EA performance in the best point
Particle(const int params)
{
ArrayResize(position, params);
ArrayResize(best, params);
ArrayResize(velocity, params);
bestValue = -DBL_MAX;
}
};
All arrays have size of parameter space dimension, that is the number of parameters being optimized. As far as our optimization implies that better performance is denoted by higher values we initialize bestValue with most minimal possible value -DBL_MAX. As you remember, EA performance can hold any of main trade performance indicators such as profit, profit factor, sharpe ratio and etc.
In addition to single particles the algorithm utilizes so called topologies or subsets of particles. They can vary, but we will use topology which simulates social groups. The group of particles will hold knowledge about best known position among all particles in the group.
{
private:
double result; // best EA performance in the group
public:
double optimum[]; // best known position in the group
Group(const int params)
{
ArrayResize(optimum, params);
ArrayInitialize(optimum, 0);
result = -DBL_MAX;
}
void assign(const double x)
{
result = x;
}
double getResult() const
{
return result;
}
bool isAssigned()
{
return result != -DBL_MAX;
}
};
Now we can start implementing the swarm algorithm itself. Of course, we pack it in a class.
{
private:
Particle *particles[];
Group *groups[];
int _size; // number of particles
int _globals; // number of groups
int _params; // number of parameters to optimize
Here we have a swarm of particles and a set of groups.
double lows[];
double steps[];
In addition we want to save optimal parameter set somewhere.
Finally we need an index array that would mark every particle as a member of specific social group.
The static array should be instantiated after the class definition.
As far as we will probably have several different constructors of the swarm, let's define a unified method for its initialization.
{
_size = size;
_globals = globals;
_params = params;
ArrayResize(particles, size);
for(int i = 0; i < size; i++)
{
particles[i] = new Particle(params);
}
ArrayResize(groups, globals);
for(int i = 0; i < globals; i++)
{
groups[i] = new Group(params);
}
ArrayResize(particle2group, size);
for(int i = 0; i < size; i++)
{
particle2group[i] = (int)MathMin(MathRand() * 1.0 / 32767 * globals, globals - 1);
}
ArrayResize(highs, params);
ArrayResize(lows, params);
ArrayResize(steps, params);
for(int i = 0; i < params; i++)
{
highs[i] = max[i];
lows[i] = min[i];
steps[i] = step[i];
}
ArrayResize(solution, params);
}
All arrays are resized to given dimensions and filled with provided data. Group membership is initialized randomly.
Swarm(const int params, const double &max[], const double &min[], const double &step[])
{
init(params * 5, (int)MathSqrt(params * 5), params, max, min, step);
}
Swarm(const int size, const int globals, const int params, const double &max[], const double &min[], const double &step[])
{
init(size, globals, params, max, min, step);
}
The first one uses some empirical rules to deduce size of the swarm and number of the groups from the number of parameters. The second allows you to specify all values explicitly.
{
for(int i = 0; i < _size; i++)
{
delete particles[i];
}
for(int i = 0; i < _globals; i++)
{
delete groups[i];
}
}
And this is the moment when the main method of the class should be coded - the method performing optimization itself.
{
Let's skip the first parameter for a while. The second one is a number of cycles which the algorithm will run. The next 3 arguments are parameters of the swarm algorithm: inertia is a coefficient to preserve particle's velocity (usually fading out, as you see from the default value 0.8), selfBoost and groupBoost define how sensitive a particle will be to requests for changing its movement in direction to best known position of the particle and its group respectively.
{
public:
virtual double calculate(const double &vector[]) = 0;
};
The method accepts an array with parameters and returns a single double value. In future we can implement a derived class in our EA, and perform virtual back-testing inside the method calculate.
First initialize result (objective function value) and coordinates in the parameter space.
ArrayInitialize(solution, 0);
Second, initialize particles. Their positions and velocities are selected randomly, and then we call the functor for every particle and save best values into groups and global solution.
{
for(int p = 0; p < _params; p++) // loop through all dimesions
{
// random placement
particles[i].position[p] = (MathRand() * 1.0 / 32767) * (highs[p] - lows[p]) + lows[p];
// the only position is the best so far
particles[i].best[p] = particles[i].position[p];
// random speed
particles[i].velocity[p] = (MathRand() * 1.0 / 32767) * 2 * (highs[p] - lows[p]) - (highs[p] - lows[p]);
}
// get the function value in the point i
particles[i].positionValue = f.calculate(particles[i].position);
// update best value of the group (if improvement is found)
if(!groups[particle2group[i]].isAssigned() || particles[i].positionValue > groups[particle2group[i]].getResult())
{
groups[particle2group[i]].assign(particles[i].positionValue);
for(int p = 0; p < _params; p++)
{
groups[particle2group[i]].optimum[p] = particles[i].position[p];
}
// update global solution (if improvement is found)
if(particles[i].positionValue > result) // global maximum
{
result = particles[i].positionValue;
for(int p = 0; p < _params; p++)
{
solution[p] = particles[i].position[p];
}
}
}
}
Then we repeat the cycle of optimization (it looks a bit lengthy but reproduces the pseudo-code almost exactly):
{
for(int i = 0; i < _size && !IsStopped(); i++) // loop through all particles
{
for(int p = 0; p < _params; p++) // update particle position and speed
{
double r1 = MathRand() * 1.0 / 32767;
double rg = MathRand() * 1.0 / 32767;
particles[i].velocity[p] = inertia * particles[i].velocity[p] + selfBoost * r1 * (particles[i].best[p] - particles[i].position[p]) + groupBoost * rg * (groups[particle2group[i]].optimum[p] - particles[i].position[p]);
particles[i].position[p] = particles[i].position[p] + particles[i].velocity[p];
// make sure to keep the particle inside the boundaries of parameter space
if(particles[i].position[p] < lows[p]) particles[i].position[p] = lows[p];
else if(particles[i].position[p] > highs[p]) particles[i].position[p] = highs[p];
}
// get the function value for the particle i
particles[i].positionValue = f.calculate(particles[i].position);
// update the particle's best value and position (if improvement is found)
if(particles[i].positionValue > particles[i].bestValue)
{
particles[i].bestValue = particles[i].positionValue;
for(int p = 0; p < _params; p++)
{
particles[i].best[p] = particles[i].position[p];
}
}
// update the group's best value and position (if improvement is found)
if(particles[i].positionValue > groups[particle2group[i]].getResult())
{
groups[particle2group[i]].assign(particles[i].positionValue);
for(int p = 0; p < _params; p++)
{
groups[particle2group[i]].optimum[p] = particles[i].position[p];
}
// update the global maximum value and solution (if improvement is found)
if(particles[i].positionValue > result)
{
result = particles[i].positionValue;
for(int p = 0; p < _params; p++)
{
solution[p] = particles[i].position[p];
}
}
}
}
}
That's all. To finalize the method we should return the result.
}
To read found solution (parameter set) from outer world we need another method.
{
ArrayResize(result, _params);
for(int p = 0; p < _params; p++)
{
result[p] = solution[p];
}
return !IsStopped();
}
Is it really all? It's too simple to be true. ;-) Actually we have a problem here.
if(steps[p] != 0)
{
particles[i].position[p] = ((int)MathRound(particles[i].position[p] / steps[p])) * steps[p];
}
This solves the problem only partially. The functor will be executed with proper parameters, but there is no guarantee that algorithm will not call it many times with the same values (due to the rounding by the step sizes). To prevent this we need to learn to identify somehow every set of values and skip repetitive calls.
It accepts CRC of a previous block of data (if any, can be 0 if only one block of data is processed; allows you to chain many blocks), the array of bytes to process and its length. The function returns 64-bit CRC.
Having this library included, we can write the following helper function to get CRC64 for an array of parameters:
#include <crc64.mqh>
template<typename T>
ulong crc64(const T &array[])
{
ulong crc = 0;
int len = ArraySize(array);
for(int i = 0; i < len; i++)
{
crc = crc64(crc, _R(array[i]).Bytes, sizeof(T));
}
return crc;
}
The next question is where to store the hashes and how to check them against every new parameter set generated by the swarm. The answer is binary tree. This is a data structure which provides very fast operations for adding new values and checking for existing ones. Actually the fastness relies on the tree property called balance. In other words the tree should be balanced to ensure the most possible quick operations. Fortunately, the fact that we'll store CRC hashes in the tree plays into our hands. Here is a short definition of a hash.
As a result, adding hashes into the binary tree makes it balanced, and automatically most efficient.
To implement this logic we need 2 classes: the node and the tree. It makes sense to use templates for both.
class TreeNode
{
private:
TreeNode *left;
TreeNode *right;
T value;
public:
TreeNode(T t)
{
value = t;
}
// adds new value into subtrees and returns false or
// returns true if t exists as value of this node or in subtrees
bool Add(T t)
{
if(t < value)
{
if(left == NULL)
{
left = new TreeNode<T>(t);
return false;
}
else
{
return left.Add(t);
}
}
else if(t > value)
{
if(right == NULL)
{
right = new TreeNode<T>(t);
return false;
}
else
{
return right.Add(t);
}
}
return true;
}
~TreeNode()
{
if(left != NULL) delete left;
if(right != NULL) delete right;
}
};
template<typename T>
class BinaryTree
{
private:
TreeNode<T> *root;
public:
bool Add(T t)
{
if(root == NULL)
{
root = new TreeNode<T>(t);
return false;
}
else
{
return root.Add(t);
}
}
~BinaryTree()
{
if(root != NULL) delete root;
}
};
I hope the algorithm is clear enough. Please note that deletion of root in the destructor will be automatically unwound to deletion of all subsidiary nodes.
{
private:
BinaryTree<ulong> index;
In the method optimize we should elaborate those parts where we move particles to new positions.
double next[];
ArrayResize(next, _params);
for(int c = 0; c < cycles && !IsStopped(); c++)
{
for(int i = 0; i < _size && !IsStopped(); i++)
{
bool alreadyProcessed;
// new placement of particles using temporary array next
for(int p = 0; p < _params; p++)
{
double r1 = MathRand() * 1.0 / 32767;
double rg = MathRand() * 1.0 / 32767;
particles[i].velocity[p] = inertia * particles[i].velocity[p] + selfBoost * r1 * (particles[i].best[p] - particles[i].position[p]) + groupBoost * rg * (groups[particle2group[i]].optimum[p] - particles[i].position[p]);
next[p] = particles[i].position[p] + particles[i].velocity[p];
if(next[p] < lows[p]) next[p] = lows[p];
else if(next[p] > highs[p]) next[p] = highs[p];
if(steps[p] != 0)
{
next[p] = ((int)MathRound(next[p] / steps[p])) * steps[p];
}
}
// check if the tree contains this parameter set and add it if not
alreadyProcessed = index.Add(crc64(next));
if(alreadyProcessed)
{
continue;
}
// apply new position to the particle
for(int p = 0; p < _params; p++)
{
particles[i].position[p] = next[p];
}
particles[i].positionValue = f.calculate(particles[i].position);
// ...
We added an auxiliary array next, where we save newly generated position and check it for uniqueness. If new position was not processed yet, we add it into the tree, copy to the particle's position and continue the procedure. If new position exists in the tree (hence has been already processed), we skip this iteration.
There exist many popular test objective functions (or benchmarks), such as "rosenbrock", "griewank", "sphere", etc. For example, for the sphere we can define the functor's calculate method as follows.
{
int dim = ArraySize(vec);
double sum = 0;
for(int i = 0; i < dim; i++) sum += pow(vec[i], 2);
return -sum; // negative for maximization
}
Please note, that conventional formulae of the test objective functions implies minimization, whereas we coded the algorithm for maximization (because we want maximal performance from EA). This is why you need to negate result in the calculation. This is important to remember if you decide to test our implementation on other popular objective functions.
Finally, all is ready to embed the fast optimization algorithm into Optimystic library for on the fly optimization of EAs. But let us address this question in the next publication.